The compare_category() compute information to examine the relationship between categorical variables.
compare_category(.data, ...)
# S3 method for class 'data.frame'
compare_category(.data, ...)Arguments
- .data
a data.frame or a
tbl_df.- ...
one or more unquoted expressions separated by commas. You can treat variable names like they are positions. Positive values select variables; negative values to drop variables. These arguments are automatically quoted and evaluated in a context where column names represent column positions. They support unquoting and splicing.
Value
An object of the class as compare based list. The information to examine the relationship between categorical variables is as follows each components.
var1 : factor. The level of the first variable to compare. 'var1' is the name of the first variable to be compared.
var2 : factor. The level of the second variable to compare. 'var2' is the name of the second variable to be compared.
n : integer. frequency by var1 and var2.
rate : double. relative frequency.
first_rate : double. relative frequency in first variable.
second_rate : double. relative frequency in second variable.
Details
It is important to understand the relationship between categorical variables in EDA. compare_category() compares relations by pair combination of all categorical variables. and return compare_category class that based list object.
Attributes of return object
Attributes of compare_category class is as follows.
variables : character. List of variables selected for comparison.
combination : matrix. It consists of pairs of variables to compare.
Examples
# \donttest{
# Generate data for the example
heartfailure2 <- heartfailure
heartfailure2[sample(seq(NROW(heartfailure2)), 5), "smoking"] <- NA
library(dplyr)
# Compare the all categorical variables
all_var <- compare_category(heartfailure2)
# Print compare_numeric class objects
all_var
#> $`anaemia vs diabetes`
#> # A tibble: 4 × 6
#> anaemia diabetes n rate var1_rate var2_rate
#> <fct> <fct> <int> <dbl> <dbl> <dbl>
#> 1 No No 98 0.328 0.576 0.563
#> 2 No Yes 72 0.241 0.424 0.576
#> 3 Yes No 76 0.254 0.589 0.437
#> 4 Yes Yes 53 0.177 0.411 0.424
#>
#> $`anaemia vs hblood_pressure`
#> # A tibble: 4 × 6
#> anaemia hblood_pressure n rate var1_rate var2_rate
#> <fct> <fct> <int> <dbl> <dbl> <dbl>
#> 1 No No 113 0.378 0.665 0.582
#> 2 No Yes 57 0.191 0.335 0.543
#> 3 Yes No 81 0.271 0.628 0.418
#> 4 Yes Yes 48 0.161 0.372 0.457
#>
#> $`anaemia vs sex`
#> # A tibble: 4 × 6
#> anaemia sex n rate var1_rate var2_rate
#> <fct> <fct> <int> <dbl> <dbl> <dbl>
#> 1 No Female 53 0.177 0.312 0.505
#> 2 No Male 117 0.391 0.688 0.603
#> 3 Yes Female 52 0.174 0.403 0.495
#> 4 Yes Male 77 0.258 0.597 0.397
#>
#> $`anaemia vs smoking`
#> # A tibble: 6 × 6
#> anaemia smoking n rate var1_rate var2_rate
#> <fct> <fct> <int> <dbl> <dbl> <dbl>
#> 1 No No 107 0.358 0.629 0.535
#> 2 No Yes 61 0.204 0.359 0.649
#> 3 No NA 2 0.00669 0.0118 0.4
#> 4 Yes No 93 0.311 0.721 0.465
#> 5 Yes Yes 33 0.110 0.256 0.351
#> 6 Yes NA 3 0.0100 0.0233 0.6
#>
#> $`anaemia vs death_event`
#> # A tibble: 4 × 6
#> anaemia death_event n rate var1_rate var2_rate
#> <fct> <fct> <int> <dbl> <dbl> <dbl>
#> 1 No No 120 0.401 0.706 0.591
#> 2 No Yes 50 0.167 0.294 0.521
#> 3 Yes No 83 0.278 0.643 0.409
#> 4 Yes Yes 46 0.154 0.357 0.479
#>
#> $`diabetes vs hblood_pressure`
#> # A tibble: 4 × 6
#> diabetes hblood_pressure n rate var1_rate var2_rate
#> <fct> <fct> <int> <dbl> <dbl> <dbl>
#> 1 No No 112 0.375 0.644 0.577
#> 2 No Yes 62 0.207 0.356 0.590
#> 3 Yes No 82 0.274 0.656 0.423
#> 4 Yes Yes 43 0.144 0.344 0.410
#>
#> $`diabetes vs sex`
#> # A tibble: 4 × 6
#> diabetes sex n rate var1_rate var2_rate
#> <fct> <fct> <int> <dbl> <dbl> <dbl>
#> 1 No Female 50 0.167 0.287 0.476
#> 2 No Male 124 0.415 0.713 0.639
#> 3 Yes Female 55 0.184 0.44 0.524
#> 4 Yes Male 70 0.234 0.56 0.361
#>
#> $`diabetes vs smoking`
#> # A tibble: 6 × 6
#> diabetes smoking n rate var1_rate var2_rate
#> <fct> <fct> <int> <dbl> <dbl> <dbl>
#> 1 No No 108 0.361 0.621 0.54
#> 2 No Yes 65 0.217 0.374 0.691
#> 3 No NA 1 0.00334 0.00575 0.2
#> 4 Yes No 92 0.308 0.736 0.46
#> 5 Yes Yes 29 0.0970 0.232 0.309
#> 6 Yes NA 4 0.0134 0.032 0.8
#>
#> $`diabetes vs death_event`
#> # A tibble: 4 × 6
#> diabetes death_event n rate var1_rate var2_rate
#> <fct> <fct> <int> <dbl> <dbl> <dbl>
#> 1 No No 118 0.395 0.678 0.581
#> 2 No Yes 56 0.187 0.322 0.583
#> 3 Yes No 85 0.284 0.68 0.419
#> 4 Yes Yes 40 0.134 0.32 0.417
#>
#> $`hblood_pressure vs sex`
#> # A tibble: 4 × 6
#> hblood_pressure sex n rate var1_rate var2_rate
#> <fct> <fct> <int> <dbl> <dbl> <dbl>
#> 1 No Female 61 0.204 0.314 0.581
#> 2 No Male 133 0.445 0.686 0.686
#> 3 Yes Female 44 0.147 0.419 0.419
#> 4 Yes Male 61 0.204 0.581 0.314
#>
#> $`hblood_pressure vs smoking`
#> # A tibble: 6 × 6
#> hblood_pressure smoking n rate var1_rate var2_rate
#> <fct> <fct> <int> <dbl> <dbl> <dbl>
#> 1 No No 127 0.425 0.655 0.635
#> 2 No Yes 64 0.214 0.330 0.681
#> 3 No NA 3 0.0100 0.0155 0.6
#> 4 Yes No 73 0.244 0.695 0.365
#> 5 Yes Yes 30 0.100 0.286 0.319
#> 6 Yes NA 2 0.00669 0.0190 0.4
#>
#> $`hblood_pressure vs death_event`
#> # A tibble: 4 × 6
#> hblood_pressure death_event n rate var1_rate var2_rate
#> <fct> <fct> <int> <dbl> <dbl> <dbl>
#> 1 No No 137 0.458 0.706 0.675
#> 2 No Yes 57 0.191 0.294 0.594
#> 3 Yes No 66 0.221 0.629 0.325
#> 4 Yes Yes 39 0.130 0.371 0.406
#>
#> $`sex vs smoking`
#> # A tibble: 6 × 6
#> sex smoking n rate var1_rate var2_rate
#> <fct> <fct> <int> <dbl> <dbl> <dbl>
#> 1 Female No 98 0.328 0.933 0.49
#> 2 Female Yes 4 0.0134 0.0381 0.0426
#> 3 Female NA 3 0.0100 0.0286 0.6
#> 4 Male No 102 0.341 0.526 0.51
#> 5 Male Yes 90 0.301 0.464 0.957
#> 6 Male NA 2 0.00669 0.0103 0.4
#>
#> $`sex vs death_event`
#> # A tibble: 4 × 6
#> sex death_event n rate var1_rate var2_rate
#> <fct> <fct> <int> <dbl> <dbl> <dbl>
#> 1 Female No 71 0.237 0.676 0.350
#> 2 Female Yes 34 0.114 0.324 0.354
#> 3 Male No 132 0.441 0.680 0.650
#> 4 Male Yes 62 0.207 0.320 0.646
#>
#> $`smoking vs death_event`
#> # A tibble: 6 × 6
#> smoking death_event n rate var1_rate var2_rate
#> <fct> <fct> <int> <dbl> <dbl> <dbl>
#> 1 No No 136 0.455 0.68 0.670
#> 2 No Yes 64 0.214 0.32 0.667
#> 3 Yes No 65 0.217 0.691 0.320
#> 4 Yes Yes 29 0.0970 0.309 0.302
#> 5 NA No 2 0.00669 0.4 0.00985
#> 6 NA Yes 3 0.0100 0.6 0.0312
#>
# Compare the categorical variables that case of joint the death_event variable
all_var %>%
"["(grep("death_event", names(all_var)))
#> $`anaemia vs death_event`
#> # A tibble: 4 × 6
#> anaemia death_event n rate var1_rate var2_rate
#> <fct> <fct> <int> <dbl> <dbl> <dbl>
#> 1 No No 120 0.401 0.706 0.591
#> 2 No Yes 50 0.167 0.294 0.521
#> 3 Yes No 83 0.278 0.643 0.409
#> 4 Yes Yes 46 0.154 0.357 0.479
#>
#> $`diabetes vs death_event`
#> # A tibble: 4 × 6
#> diabetes death_event n rate var1_rate var2_rate
#> <fct> <fct> <int> <dbl> <dbl> <dbl>
#> 1 No No 118 0.395 0.678 0.581
#> 2 No Yes 56 0.187 0.322 0.583
#> 3 Yes No 85 0.284 0.68 0.419
#> 4 Yes Yes 40 0.134 0.32 0.417
#>
#> $`hblood_pressure vs death_event`
#> # A tibble: 4 × 6
#> hblood_pressure death_event n rate var1_rate var2_rate
#> <fct> <fct> <int> <dbl> <dbl> <dbl>
#> 1 No No 137 0.458 0.706 0.675
#> 2 No Yes 57 0.191 0.294 0.594
#> 3 Yes No 66 0.221 0.629 0.325
#> 4 Yes Yes 39 0.130 0.371 0.406
#>
#> $`sex vs death_event`
#> # A tibble: 4 × 6
#> sex death_event n rate var1_rate var2_rate
#> <fct> <fct> <int> <dbl> <dbl> <dbl>
#> 1 Female No 71 0.237 0.676 0.350
#> 2 Female Yes 34 0.114 0.324 0.354
#> 3 Male No 132 0.441 0.680 0.650
#> 4 Male Yes 62 0.207 0.320 0.646
#>
#> $`smoking vs death_event`
#> # A tibble: 6 × 6
#> smoking death_event n rate var1_rate var2_rate
#> <fct> <fct> <int> <dbl> <dbl> <dbl>
#> 1 No No 136 0.455 0.68 0.670
#> 2 No Yes 64 0.214 0.32 0.667
#> 3 Yes No 65 0.217 0.691 0.320
#> 4 Yes Yes 29 0.0970 0.309 0.302
#> 5 NA No 2 0.00669 0.4 0.00985
#> 6 NA Yes 3 0.0100 0.6 0.0312
#>
# Compare the two categorical variables
two_var <- compare_category(heartfailure2, smoking, death_event)
# Print compare_category class objects
two_var
#> $`smoking vs death_event`
#> # A tibble: 6 × 6
#> smoking death_event n rate var1_rate var2_rate
#> <fct> <fct> <int> <dbl> <dbl> <dbl>
#> 1 No No 136 0.455 0.68 0.670
#> 2 No Yes 64 0.214 0.32 0.667
#> 3 Yes No 65 0.217 0.691 0.320
#> 4 Yes Yes 29 0.0970 0.309 0.302
#> 5 NA No 2 0.00669 0.4 0.00985
#> 6 NA Yes 3 0.0100 0.6 0.0312
#>
# Filtering the case of smoking included NA
two_var %>%
"[["(1) %>%
filter(!is.na(smoking))
#> # A tibble: 4 × 6
#> smoking death_event n rate var1_rate var2_rate
#> <fct> <fct> <int> <dbl> <dbl> <dbl>
#> 1 No No 136 0.455 0.68 0.670
#> 2 No Yes 64 0.214 0.32 0.667
#> 3 Yes No 65 0.217 0.691 0.320
#> 4 Yes Yes 29 0.0970 0.309 0.302
# Summary the all case : Return a invisible copy of an object.
stat <- summary(all_var)
#> ── Contingency tables ──────────────────────────── Number of table is 15 ──
#> $`anaemia vs diabetes`
#> diabetes
#> anaemia No Yes
#> No 98 72
#> Yes 76 53
#>
#> $`anaemia vs hblood_pressure`
#> hblood_pressure
#> anaemia No Yes
#> No 113 57
#> Yes 81 48
#>
#> $`anaemia vs sex`
#> sex
#> anaemia Female Male
#> No 53 117
#> Yes 52 77
#>
#> $`anaemia vs smoking`
#> smoking
#> anaemia No Yes
#> No 107 61
#> Yes 93 33
#>
#> $`anaemia vs death_event`
#> death_event
#> anaemia No Yes
#> No 120 50
#> Yes 83 46
#>
#> $`diabetes vs hblood_pressure`
#> hblood_pressure
#> diabetes No Yes
#> No 112 62
#> Yes 82 43
#>
#> $`diabetes vs sex`
#> sex
#> diabetes Female Male
#> No 50 124
#> Yes 55 70
#>
#> $`diabetes vs smoking`
#> smoking
#> diabetes No Yes
#> No 108 65
#> Yes 92 29
#>
#> $`diabetes vs death_event`
#> death_event
#> diabetes No Yes
#> No 118 56
#> Yes 85 40
#>
#> $`hblood_pressure vs sex`
#> sex
#> hblood_pressure Female Male
#> No 61 133
#> Yes 44 61
#>
#> $`hblood_pressure vs smoking`
#> smoking
#> hblood_pressure No Yes
#> No 127 64
#> Yes 73 30
#>
#> $`hblood_pressure vs death_event`
#> death_event
#> hblood_pressure No Yes
#> No 137 57
#> Yes 66 39
#>
#> $`sex vs smoking`
#> smoking
#> sex No Yes
#> Female 98 4
#> Male 102 90
#>
#> $`sex vs death_event`
#> death_event
#> sex No Yes
#> Female 71 34
#> Male 132 62
#>
#> $`smoking vs death_event`
#> death_event
#> smoking No Yes
#> No 136 64
#> Yes 65 29
#>
#> ── Relative contingency tables ─────────────────── Number of table is 15 ──
#> $`anaemia vs diabetes`
#> diabetes
#> anaemia No Yes
#> No 0.3277592 0.2408027
#> Yes 0.2541806 0.1772575
#>
#> $`anaemia vs hblood_pressure`
#> hblood_pressure
#> anaemia No Yes
#> No 0.3779264 0.1906355
#> Yes 0.2709030 0.1605351
#>
#> $`anaemia vs sex`
#> sex
#> anaemia Female Male
#> No 0.1772575 0.3913043
#> Yes 0.1739130 0.2575251
#>
#> $`anaemia vs smoking`
#> smoking
#> anaemia No Yes
#> No 0.3639456 0.2074830
#> Yes 0.3163265 0.1122449
#>
#> $`anaemia vs death_event`
#> death_event
#> anaemia No Yes
#> No 0.4013378 0.1672241
#> Yes 0.2775920 0.1538462
#>
#> $`diabetes vs hblood_pressure`
#> hblood_pressure
#> diabetes No Yes
#> No 0.3745819 0.2073579
#> Yes 0.2742475 0.1438127
#>
#> $`diabetes vs sex`
#> sex
#> diabetes Female Male
#> No 0.1672241 0.4147157
#> Yes 0.1839465 0.2341137
#>
#> $`diabetes vs smoking`
#> smoking
#> diabetes No Yes
#> No 0.36734694 0.22108844
#> Yes 0.31292517 0.09863946
#>
#> $`diabetes vs death_event`
#> death_event
#> diabetes No Yes
#> No 0.3946488 0.1872910
#> Yes 0.2842809 0.1337793
#>
#> $`hblood_pressure vs sex`
#> sex
#> hblood_pressure Female Male
#> No 0.2040134 0.4448161
#> Yes 0.1471572 0.2040134
#>
#> $`hblood_pressure vs smoking`
#> smoking
#> hblood_pressure No Yes
#> No 0.4319728 0.2176871
#> Yes 0.2482993 0.1020408
#>
#> $`hblood_pressure vs death_event`
#> death_event
#> hblood_pressure No Yes
#> No 0.4581940 0.1906355
#> Yes 0.2207358 0.1304348
#>
#> $`sex vs smoking`
#> smoking
#> sex No Yes
#> Female 0.33333333 0.01360544
#> Male 0.34693878 0.30612245
#>
#> $`sex vs death_event`
#> death_event
#> sex No Yes
#> Female 0.2374582 0.1137124
#> Male 0.4414716 0.2073579
#>
#> $`smoking vs death_event`
#> death_event
#> smoking No Yes
#> No 0.46258503 0.21768707
#> Yes 0.22108844 0.09863946
#>
#> ── Chi-squared contingency table tests ─────────── Number of table is 15 ──
#> variable_1 variable_2 statistic p.value df
#> 1 anaemia diabetes 1.035093e-02 9.189634e-01 1
#> 2 anaemia hblood_pressure 2.893564e-01 5.906333e-01 1
#> 3 anaemia sex 2.299464e+00 1.294186e-01 1
#> 4 anaemia smoking 2.940326e+00 8.639331e-02 1
#> 5 anaemia death_event 1.042175e+00 3.073161e-01 1
#> 6 diabetes hblood_pressure 9.476710e-03 9.224497e-01 1
#> 7 diabetes sex 6.783853e+00 9.198613e-03 1
#> 8 diabetes smoking 5.450134e+00 1.956683e-02 1
#> 9 diabetes death_event 2.161684e-30 1.000000e+00 1
#> 10 hblood_pressure sex 2.829289e+00 9.255934e-02 1
#> 11 hblood_pressure smoking 4.063787e-01 5.238133e-01 1
#> 12 hblood_pressure death_event 1.543461e+00 2.141034e-01 1
#> 13 sex smoking 5.454737e+01 1.517458e-13 1
#> 14 sex death_event 0.000000e+00 1.000000e+00 1
#> 15 smoking death_event 3.982985e-03 9.496782e-01 1
# Summary by returned objects
stat
#> $table
#> $table$`anaemia vs diabetes`
#> diabetes
#> anaemia No Yes
#> No 98 72
#> Yes 76 53
#>
#> $table$`anaemia vs hblood_pressure`
#> hblood_pressure
#> anaemia No Yes
#> No 113 57
#> Yes 81 48
#>
#> $table$`anaemia vs sex`
#> sex
#> anaemia Female Male
#> No 53 117
#> Yes 52 77
#>
#> $table$`anaemia vs smoking`
#> smoking
#> anaemia No Yes
#> No 107 61
#> Yes 93 33
#>
#> $table$`anaemia vs death_event`
#> death_event
#> anaemia No Yes
#> No 120 50
#> Yes 83 46
#>
#> $table$`diabetes vs hblood_pressure`
#> hblood_pressure
#> diabetes No Yes
#> No 112 62
#> Yes 82 43
#>
#> $table$`diabetes vs sex`
#> sex
#> diabetes Female Male
#> No 50 124
#> Yes 55 70
#>
#> $table$`diabetes vs smoking`
#> smoking
#> diabetes No Yes
#> No 108 65
#> Yes 92 29
#>
#> $table$`diabetes vs death_event`
#> death_event
#> diabetes No Yes
#> No 118 56
#> Yes 85 40
#>
#> $table$`hblood_pressure vs sex`
#> sex
#> hblood_pressure Female Male
#> No 61 133
#> Yes 44 61
#>
#> $table$`hblood_pressure vs smoking`
#> smoking
#> hblood_pressure No Yes
#> No 127 64
#> Yes 73 30
#>
#> $table$`hblood_pressure vs death_event`
#> death_event
#> hblood_pressure No Yes
#> No 137 57
#> Yes 66 39
#>
#> $table$`sex vs smoking`
#> smoking
#> sex No Yes
#> Female 98 4
#> Male 102 90
#>
#> $table$`sex vs death_event`
#> death_event
#> sex No Yes
#> Female 71 34
#> Male 132 62
#>
#> $table$`smoking vs death_event`
#> death_event
#> smoking No Yes
#> No 136 64
#> Yes 65 29
#>
#>
#> $relative
#> $relative$`anaemia vs diabetes`
#> diabetes
#> anaemia No Yes
#> No 0.3277592 0.2408027
#> Yes 0.2541806 0.1772575
#>
#> $relative$`anaemia vs hblood_pressure`
#> hblood_pressure
#> anaemia No Yes
#> No 0.3779264 0.1906355
#> Yes 0.2709030 0.1605351
#>
#> $relative$`anaemia vs sex`
#> sex
#> anaemia Female Male
#> No 0.1772575 0.3913043
#> Yes 0.1739130 0.2575251
#>
#> $relative$`anaemia vs smoking`
#> smoking
#> anaemia No Yes
#> No 0.3639456 0.2074830
#> Yes 0.3163265 0.1122449
#>
#> $relative$`anaemia vs death_event`
#> death_event
#> anaemia No Yes
#> No 0.4013378 0.1672241
#> Yes 0.2775920 0.1538462
#>
#> $relative$`diabetes vs hblood_pressure`
#> hblood_pressure
#> diabetes No Yes
#> No 0.3745819 0.2073579
#> Yes 0.2742475 0.1438127
#>
#> $relative$`diabetes vs sex`
#> sex
#> diabetes Female Male
#> No 0.1672241 0.4147157
#> Yes 0.1839465 0.2341137
#>
#> $relative$`diabetes vs smoking`
#> smoking
#> diabetes No Yes
#> No 0.36734694 0.22108844
#> Yes 0.31292517 0.09863946
#>
#> $relative$`diabetes vs death_event`
#> death_event
#> diabetes No Yes
#> No 0.3946488 0.1872910
#> Yes 0.2842809 0.1337793
#>
#> $relative$`hblood_pressure vs sex`
#> sex
#> hblood_pressure Female Male
#> No 0.2040134 0.4448161
#> Yes 0.1471572 0.2040134
#>
#> $relative$`hblood_pressure vs smoking`
#> smoking
#> hblood_pressure No Yes
#> No 0.4319728 0.2176871
#> Yes 0.2482993 0.1020408
#>
#> $relative$`hblood_pressure vs death_event`
#> death_event
#> hblood_pressure No Yes
#> No 0.4581940 0.1906355
#> Yes 0.2207358 0.1304348
#>
#> $relative$`sex vs smoking`
#> smoking
#> sex No Yes
#> Female 0.33333333 0.01360544
#> Male 0.34693878 0.30612245
#>
#> $relative$`sex vs death_event`
#> death_event
#> sex No Yes
#> Female 0.2374582 0.1137124
#> Male 0.4414716 0.2073579
#>
#> $relative$`smoking vs death_event`
#> death_event
#> smoking No Yes
#> No 0.46258503 0.21768707
#> Yes 0.22108844 0.09863946
#>
#>
#> $chisq
#> variable_1 variable_2 statistic p.value df
#> 1 anaemia diabetes 1.035093e-02 9.189634e-01 1
#> 2 anaemia hblood_pressure 2.893564e-01 5.906333e-01 1
#> 3 anaemia sex 2.299464e+00 1.294186e-01 1
#> 4 anaemia smoking 2.940326e+00 8.639331e-02 1
#> 5 anaemia death_event 1.042175e+00 3.073161e-01 1
#> 6 diabetes hblood_pressure 9.476710e-03 9.224497e-01 1
#> 7 diabetes sex 6.783853e+00 9.198613e-03 1
#> 8 diabetes smoking 5.450134e+00 1.956683e-02 1
#> 9 diabetes death_event 2.161684e-30 1.000000e+00 1
#> 10 hblood_pressure sex 2.829289e+00 9.255934e-02 1
#> 11 hblood_pressure smoking 4.063787e-01 5.238133e-01 1
#> 12 hblood_pressure death_event 1.543461e+00 2.141034e-01 1
#> 13 sex smoking 5.454737e+01 1.517458e-13 1
#> 14 sex death_event 0.000000e+00 1.000000e+00 1
#> 15 smoking death_event 3.982985e-03 9.496782e-01 1
#>
# component of table
stat$table
#> $`anaemia vs diabetes`
#> diabetes
#> anaemia No Yes
#> No 98 72
#> Yes 76 53
#>
#> $`anaemia vs hblood_pressure`
#> hblood_pressure
#> anaemia No Yes
#> No 113 57
#> Yes 81 48
#>
#> $`anaemia vs sex`
#> sex
#> anaemia Female Male
#> No 53 117
#> Yes 52 77
#>
#> $`anaemia vs smoking`
#> smoking
#> anaemia No Yes
#> No 107 61
#> Yes 93 33
#>
#> $`anaemia vs death_event`
#> death_event
#> anaemia No Yes
#> No 120 50
#> Yes 83 46
#>
#> $`diabetes vs hblood_pressure`
#> hblood_pressure
#> diabetes No Yes
#> No 112 62
#> Yes 82 43
#>
#> $`diabetes vs sex`
#> sex
#> diabetes Female Male
#> No 50 124
#> Yes 55 70
#>
#> $`diabetes vs smoking`
#> smoking
#> diabetes No Yes
#> No 108 65
#> Yes 92 29
#>
#> $`diabetes vs death_event`
#> death_event
#> diabetes No Yes
#> No 118 56
#> Yes 85 40
#>
#> $`hblood_pressure vs sex`
#> sex
#> hblood_pressure Female Male
#> No 61 133
#> Yes 44 61
#>
#> $`hblood_pressure vs smoking`
#> smoking
#> hblood_pressure No Yes
#> No 127 64
#> Yes 73 30
#>
#> $`hblood_pressure vs death_event`
#> death_event
#> hblood_pressure No Yes
#> No 137 57
#> Yes 66 39
#>
#> $`sex vs smoking`
#> smoking
#> sex No Yes
#> Female 98 4
#> Male 102 90
#>
#> $`sex vs death_event`
#> death_event
#> sex No Yes
#> Female 71 34
#> Male 132 62
#>
#> $`smoking vs death_event`
#> death_event
#> smoking No Yes
#> No 136 64
#> Yes 65 29
#>
# component of chi-square test
stat$chisq
#> variable_1 variable_2 statistic p.value df
#> 1 anaemia diabetes 1.035093e-02 9.189634e-01 1
#> 2 anaemia hblood_pressure 2.893564e-01 5.906333e-01 1
#> 3 anaemia sex 2.299464e+00 1.294186e-01 1
#> 4 anaemia smoking 2.940326e+00 8.639331e-02 1
#> 5 anaemia death_event 1.042175e+00 3.073161e-01 1
#> 6 diabetes hblood_pressure 9.476710e-03 9.224497e-01 1
#> 7 diabetes sex 6.783853e+00 9.198613e-03 1
#> 8 diabetes smoking 5.450134e+00 1.956683e-02 1
#> 9 diabetes death_event 2.161684e-30 1.000000e+00 1
#> 10 hblood_pressure sex 2.829289e+00 9.255934e-02 1
#> 11 hblood_pressure smoking 4.063787e-01 5.238133e-01 1
#> 12 hblood_pressure death_event 1.543461e+00 2.141034e-01 1
#> 13 sex smoking 5.454737e+01 1.517458e-13 1
#> 14 sex death_event 0.000000e+00 1.000000e+00 1
#> 15 smoking death_event 3.982985e-03 9.496782e-01 1
# component of chi-square test
summary(all_var, "chisq")
#> ── Chi-squared contingency table tests ─────────── Number of table is 15 ──
#> variable_1 variable_2 statistic p.value df
#> 1 anaemia diabetes 1.035093e-02 9.189634e-01 1
#> 2 anaemia hblood_pressure 2.893564e-01 5.906333e-01 1
#> 3 anaemia sex 2.299464e+00 1.294186e-01 1
#> 4 anaemia smoking 2.940326e+00 8.639331e-02 1
#> 5 anaemia death_event 1.042175e+00 3.073161e-01 1
#> 6 diabetes hblood_pressure 9.476710e-03 9.224497e-01 1
#> 7 diabetes sex 6.783853e+00 9.198613e-03 1
#> 8 diabetes smoking 5.450134e+00 1.956683e-02 1
#> 9 diabetes death_event 2.161684e-30 1.000000e+00 1
#> 10 hblood_pressure sex 2.829289e+00 9.255934e-02 1
#> 11 hblood_pressure smoking 4.063787e-01 5.238133e-01 1
#> 12 hblood_pressure death_event 1.543461e+00 2.141034e-01 1
#> 13 sex smoking 5.454737e+01 1.517458e-13 1
#> 14 sex death_event 0.000000e+00 1.000000e+00 1
#> 15 smoking death_event 3.982985e-03 9.496782e-01 1
# component of chi-square test (first, third case)
summary(all_var, "chisq", pos = c(1, 3))
#> ── Chi-squared contingency table tests ──────────── Number of table is 2 ──
#> variable_1 variable_2 statistic p.value df
#> 1 anaemia diabetes 0.01035093 0.9189634 1
#> 2 anaemia sex 2.29946450 0.1294186 1
# component of relative frequency table
summary(all_var, "relative")
#> ── Relative contingency tables ─────────────────── Number of table is 15 ──
#> $`anaemia vs diabetes`
#> diabetes
#> anaemia No Yes
#> No 0.3277592 0.2408027
#> Yes 0.2541806 0.1772575
#>
#> $`anaemia vs hblood_pressure`
#> hblood_pressure
#> anaemia No Yes
#> No 0.3779264 0.1906355
#> Yes 0.2709030 0.1605351
#>
#> $`anaemia vs sex`
#> sex
#> anaemia Female Male
#> No 0.1772575 0.3913043
#> Yes 0.1739130 0.2575251
#>
#> $`anaemia vs smoking`
#> smoking
#> anaemia No Yes
#> No 0.3639456 0.2074830
#> Yes 0.3163265 0.1122449
#>
#> $`anaemia vs death_event`
#> death_event
#> anaemia No Yes
#> No 0.4013378 0.1672241
#> Yes 0.2775920 0.1538462
#>
#> $`diabetes vs hblood_pressure`
#> hblood_pressure
#> diabetes No Yes
#> No 0.3745819 0.2073579
#> Yes 0.2742475 0.1438127
#>
#> $`diabetes vs sex`
#> sex
#> diabetes Female Male
#> No 0.1672241 0.4147157
#> Yes 0.1839465 0.2341137
#>
#> $`diabetes vs smoking`
#> smoking
#> diabetes No Yes
#> No 0.36734694 0.22108844
#> Yes 0.31292517 0.09863946
#>
#> $`diabetes vs death_event`
#> death_event
#> diabetes No Yes
#> No 0.3946488 0.1872910
#> Yes 0.2842809 0.1337793
#>
#> $`hblood_pressure vs sex`
#> sex
#> hblood_pressure Female Male
#> No 0.2040134 0.4448161
#> Yes 0.1471572 0.2040134
#>
#> $`hblood_pressure vs smoking`
#> smoking
#> hblood_pressure No Yes
#> No 0.4319728 0.2176871
#> Yes 0.2482993 0.1020408
#>
#> $`hblood_pressure vs death_event`
#> death_event
#> hblood_pressure No Yes
#> No 0.4581940 0.1906355
#> Yes 0.2207358 0.1304348
#>
#> $`sex vs smoking`
#> smoking
#> sex No Yes
#> Female 0.33333333 0.01360544
#> Male 0.34693878 0.30612245
#>
#> $`sex vs death_event`
#> death_event
#> sex No Yes
#> Female 0.2374582 0.1137124
#> Male 0.4414716 0.2073579
#>
#> $`smoking vs death_event`
#> death_event
#> smoking No Yes
#> No 0.46258503 0.21768707
#> Yes 0.22108844 0.09863946
#>
# component of table without missing values
summary(all_var, "table", na.rm = TRUE)
#> ── Contingency tables ──────────────────────────── Number of table is 15 ──
#> $`anaemia vs diabetes`
#> diabetes
#> anaemia No Yes
#> No 98 72
#> Yes 76 53
#>
#> $`anaemia vs hblood_pressure`
#> hblood_pressure
#> anaemia No Yes
#> No 113 57
#> Yes 81 48
#>
#> $`anaemia vs sex`
#> sex
#> anaemia Female Male
#> No 53 117
#> Yes 52 77
#>
#> $`anaemia vs smoking`
#> smoking
#> anaemia No Yes
#> No 107 61
#> Yes 93 33
#>
#> $`anaemia vs death_event`
#> death_event
#> anaemia No Yes
#> No 120 50
#> Yes 83 46
#>
#> $`diabetes vs hblood_pressure`
#> hblood_pressure
#> diabetes No Yes
#> No 112 62
#> Yes 82 43
#>
#> $`diabetes vs sex`
#> sex
#> diabetes Female Male
#> No 50 124
#> Yes 55 70
#>
#> $`diabetes vs smoking`
#> smoking
#> diabetes No Yes
#> No 108 65
#> Yes 92 29
#>
#> $`diabetes vs death_event`
#> death_event
#> diabetes No Yes
#> No 118 56
#> Yes 85 40
#>
#> $`hblood_pressure vs sex`
#> sex
#> hblood_pressure Female Male
#> No 61 133
#> Yes 44 61
#>
#> $`hblood_pressure vs smoking`
#> smoking
#> hblood_pressure No Yes
#> No 127 64
#> Yes 73 30
#>
#> $`hblood_pressure vs death_event`
#> death_event
#> hblood_pressure No Yes
#> No 137 57
#> Yes 66 39
#>
#> $`sex vs smoking`
#> smoking
#> sex No Yes
#> Female 98 4
#> Male 102 90
#>
#> $`sex vs death_event`
#> death_event
#> sex No Yes
#> Female 71 34
#> Male 132 62
#>
#> $`smoking vs death_event`
#> death_event
#> smoking No Yes
#> No 136 64
#> Yes 65 29
#>
# component of table include marginal value
margin <- summary(all_var, "table", marginal = TRUE)
#> ── Contingency tables ──────────────────────────── Number of table is 15 ──
#> $`anaemia vs diabetes`
#> diabetes
#> anaemia No Yes <Total>
#> No 98 72 170
#> Yes 76 53 129
#> <Total> 174 125 299
#>
#> $`anaemia vs hblood_pressure`
#> hblood_pressure
#> anaemia No Yes <Total>
#> No 113 57 170
#> Yes 81 48 129
#> <Total> 194 105 299
#>
#> $`anaemia vs sex`
#> sex
#> anaemia Female Male <Total>
#> No 53 117 170
#> Yes 52 77 129
#> <Total> 105 194 299
#>
#> $`anaemia vs smoking`
#> smoking
#> anaemia No Yes <Total>
#> No 107 61 168
#> Yes 93 33 126
#> <Total> 200 94 294
#>
#> $`anaemia vs death_event`
#> death_event
#> anaemia No Yes <Total>
#> No 120 50 170
#> Yes 83 46 129
#> <Total> 203 96 299
#>
#> $`diabetes vs hblood_pressure`
#> hblood_pressure
#> diabetes No Yes <Total>
#> No 112 62 174
#> Yes 82 43 125
#> <Total> 194 105 299
#>
#> $`diabetes vs sex`
#> sex
#> diabetes Female Male <Total>
#> No 50 124 174
#> Yes 55 70 125
#> <Total> 105 194 299
#>
#> $`diabetes vs smoking`
#> smoking
#> diabetes No Yes <Total>
#> No 108 65 173
#> Yes 92 29 121
#> <Total> 200 94 294
#>
#> $`diabetes vs death_event`
#> death_event
#> diabetes No Yes <Total>
#> No 118 56 174
#> Yes 85 40 125
#> <Total> 203 96 299
#>
#> $`hblood_pressure vs sex`
#> sex
#> hblood_pressure Female Male <Total>
#> No 61 133 194
#> Yes 44 61 105
#> <Total> 105 194 299
#>
#> $`hblood_pressure vs smoking`
#> smoking
#> hblood_pressure No Yes <Total>
#> No 127 64 191
#> Yes 73 30 103
#> <Total> 200 94 294
#>
#> $`hblood_pressure vs death_event`
#> death_event
#> hblood_pressure No Yes <Total>
#> No 137 57 194
#> Yes 66 39 105
#> <Total> 203 96 299
#>
#> $`sex vs smoking`
#> smoking
#> sex No Yes <Total>
#> Female 98 4 102
#> Male 102 90 192
#> <Total> 200 94 294
#>
#> $`sex vs death_event`
#> death_event
#> sex No Yes <Total>
#> Female 71 34 105
#> Male 132 62 194
#> <Total> 203 96 299
#>
#> $`smoking vs death_event`
#> death_event
#> smoking No Yes <Total>
#> No 136 64 200
#> Yes 65 29 94
#> <Total> 201 93 294
#>
margin
#> $`anaemia vs diabetes`
#> diabetes
#> anaemia No Yes <Total>
#> No 98 72 170
#> Yes 76 53 129
#> <Total> 174 125 299
#>
#> $`anaemia vs hblood_pressure`
#> hblood_pressure
#> anaemia No Yes <Total>
#> No 113 57 170
#> Yes 81 48 129
#> <Total> 194 105 299
#>
#> $`anaemia vs sex`
#> sex
#> anaemia Female Male <Total>
#> No 53 117 170
#> Yes 52 77 129
#> <Total> 105 194 299
#>
#> $`anaemia vs smoking`
#> smoking
#> anaemia No Yes <Total>
#> No 107 61 168
#> Yes 93 33 126
#> <Total> 200 94 294
#>
#> $`anaemia vs death_event`
#> death_event
#> anaemia No Yes <Total>
#> No 120 50 170
#> Yes 83 46 129
#> <Total> 203 96 299
#>
#> $`diabetes vs hblood_pressure`
#> hblood_pressure
#> diabetes No Yes <Total>
#> No 112 62 174
#> Yes 82 43 125
#> <Total> 194 105 299
#>
#> $`diabetes vs sex`
#> sex
#> diabetes Female Male <Total>
#> No 50 124 174
#> Yes 55 70 125
#> <Total> 105 194 299
#>
#> $`diabetes vs smoking`
#> smoking
#> diabetes No Yes <Total>
#> No 108 65 173
#> Yes 92 29 121
#> <Total> 200 94 294
#>
#> $`diabetes vs death_event`
#> death_event
#> diabetes No Yes <Total>
#> No 118 56 174
#> Yes 85 40 125
#> <Total> 203 96 299
#>
#> $`hblood_pressure vs sex`
#> sex
#> hblood_pressure Female Male <Total>
#> No 61 133 194
#> Yes 44 61 105
#> <Total> 105 194 299
#>
#> $`hblood_pressure vs smoking`
#> smoking
#> hblood_pressure No Yes <Total>
#> No 127 64 191
#> Yes 73 30 103
#> <Total> 200 94 294
#>
#> $`hblood_pressure vs death_event`
#> death_event
#> hblood_pressure No Yes <Total>
#> No 137 57 194
#> Yes 66 39 105
#> <Total> 203 96 299
#>
#> $`sex vs smoking`
#> smoking
#> sex No Yes <Total>
#> Female 98 4 102
#> Male 102 90 192
#> <Total> 200 94 294
#>
#> $`sex vs death_event`
#> death_event
#> sex No Yes <Total>
#> Female 71 34 105
#> Male 132 62 194
#> <Total> 203 96 299
#>
#> $`smoking vs death_event`
#> death_event
#> smoking No Yes <Total>
#> No 136 64 200
#> Yes 65 29 94
#> <Total> 201 93 294
#>
# component of chi-square test
summary(two_var, method = "chisq")
#> ── Chi-squared contingency table tests ──────────── Number of table is 1 ──
#> variable_1 variable_2 statistic p.value df
#> 1 smoking death_event 0.003982985 0.9496782 1
# verbose is FALSE
summary(all_var, "chisq", verbose = FALSE)
#> variable_1 variable_2 statistic p.value df
#> 1 anaemia diabetes 1.035093e-02 9.189634e-01 1
#> 2 anaemia hblood_pressure 2.893564e-01 5.906333e-01 1
#> 3 anaemia sex 2.299464e+00 1.294186e-01 1
#> 4 anaemia smoking 2.940326e+00 8.639331e-02 1
#> 5 anaemia death_event 1.042175e+00 3.073161e-01 1
#> 6 diabetes hblood_pressure 9.476710e-03 9.224497e-01 1
#> 7 diabetes sex 6.783853e+00 9.198613e-03 1
#> 8 diabetes smoking 5.450134e+00 1.956683e-02 1
#> 9 diabetes death_event 2.161684e-30 1.000000e+00 1
#> 10 hblood_pressure sex 2.829289e+00 9.255934e-02 1
#> 11 hblood_pressure smoking 4.063787e-01 5.238133e-01 1
#> 12 hblood_pressure death_event 1.543461e+00 2.141034e-01 1
#> 13 sex smoking 5.454737e+01 1.517458e-13 1
#> 14 sex death_event 0.000000e+00 1.000000e+00 1
#> 15 smoking death_event 3.982985e-03 9.496782e-01 1
#' # Using pipes & dplyr -------------------------
# If you want to use dplyr, set verbose to FALSE
summary(all_var, "chisq", verbose = FALSE) %>%
filter(p.value < 0.26)
#> variable_1 variable_2 statistic p.value df
#> 1 anaemia sex 2.299464 1.294186e-01 1
#> 2 anaemia smoking 2.940326 8.639331e-02 1
#> 3 diabetes sex 6.783853 9.198613e-03 1
#> 4 diabetes smoking 5.450134 1.956683e-02 1
#> 5 hblood_pressure sex 2.829289 9.255934e-02 1
#> 6 hblood_pressure death_event 1.543461 2.141034e-01 1
#> 7 sex smoking 54.547368 1.517458e-13 1
# Extract component from list by index
summary(all_var, "table", na.rm = TRUE, verbose = FALSE) %>%
"[["(1)
#> diabetes
#> anaemia No Yes
#> No 98 72
#> Yes 76 53
# Extract component from list by name
summary(all_var, "table", na.rm = TRUE, verbose = FALSE) %>%
"[["("smoking vs death_event")
#> death_event
#> smoking No Yes
#> No 136 64
#> Yes 65 29
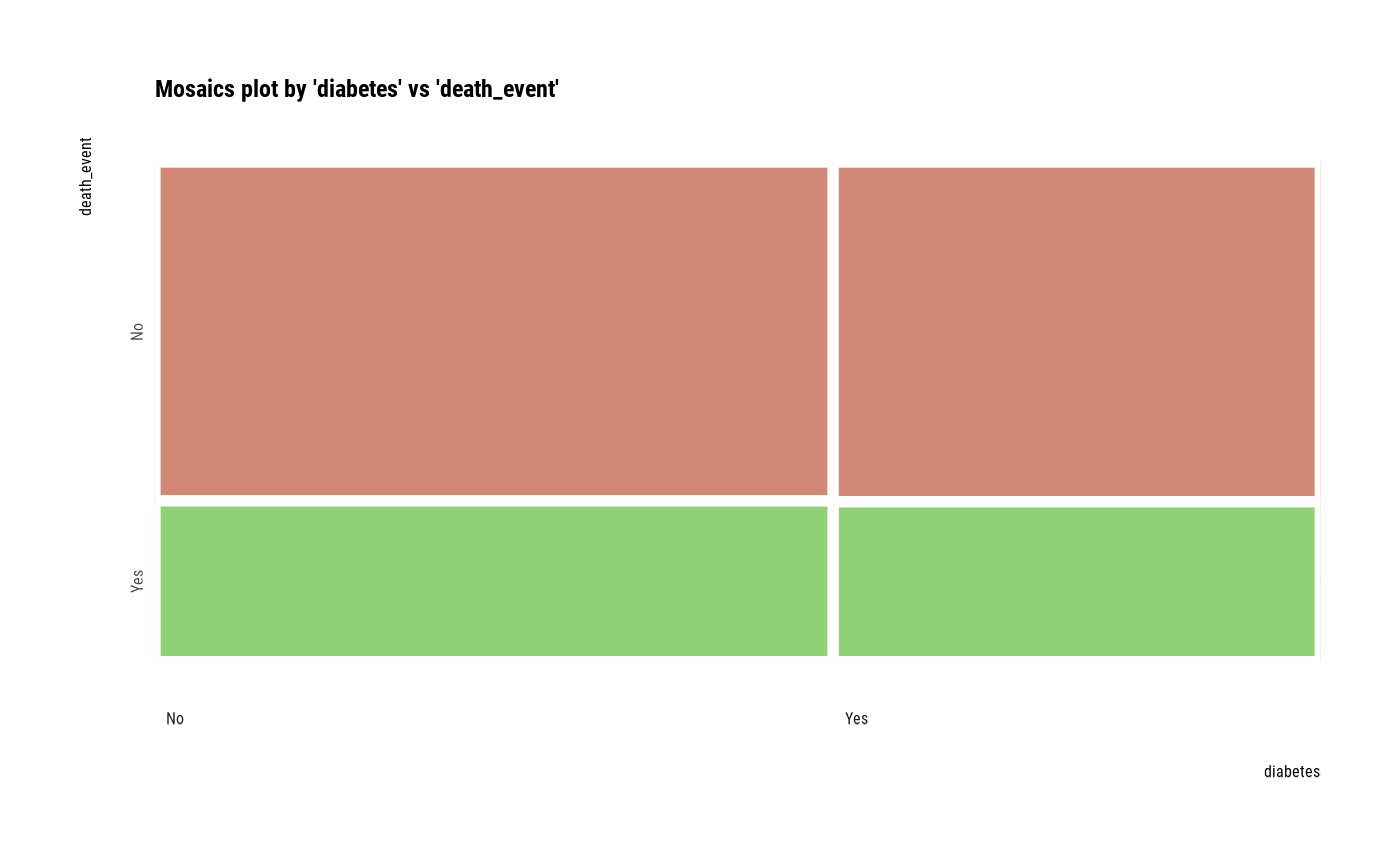
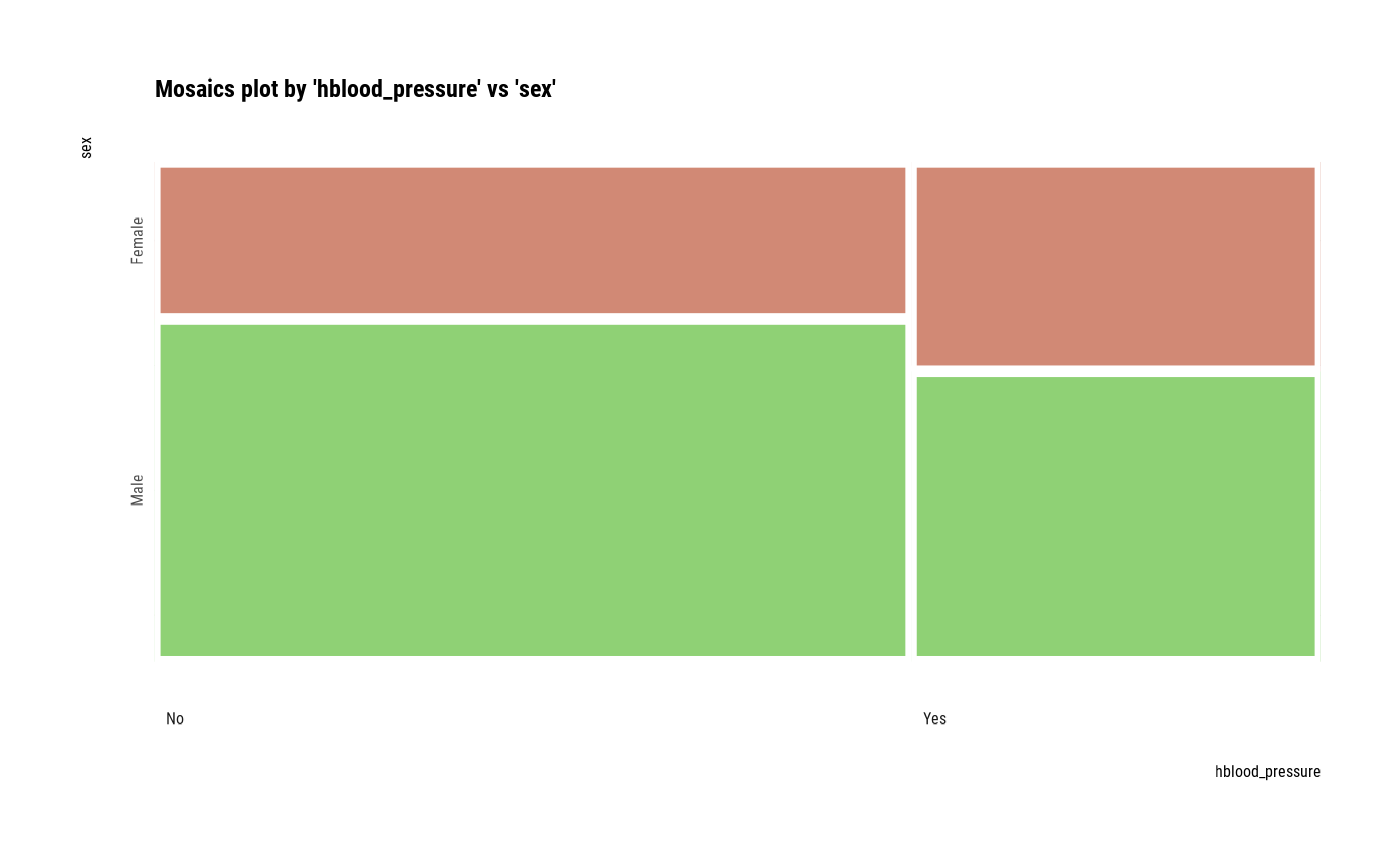
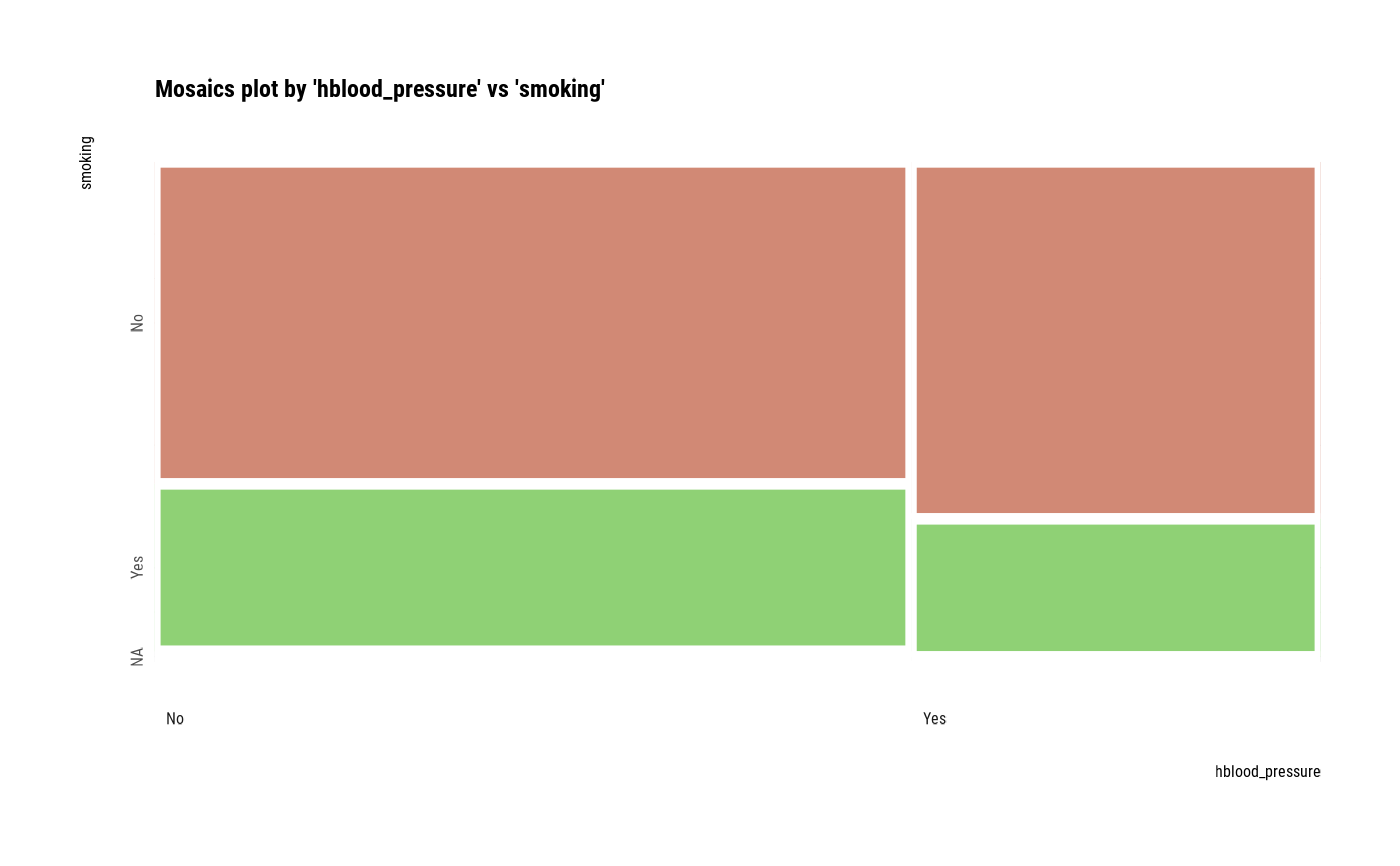
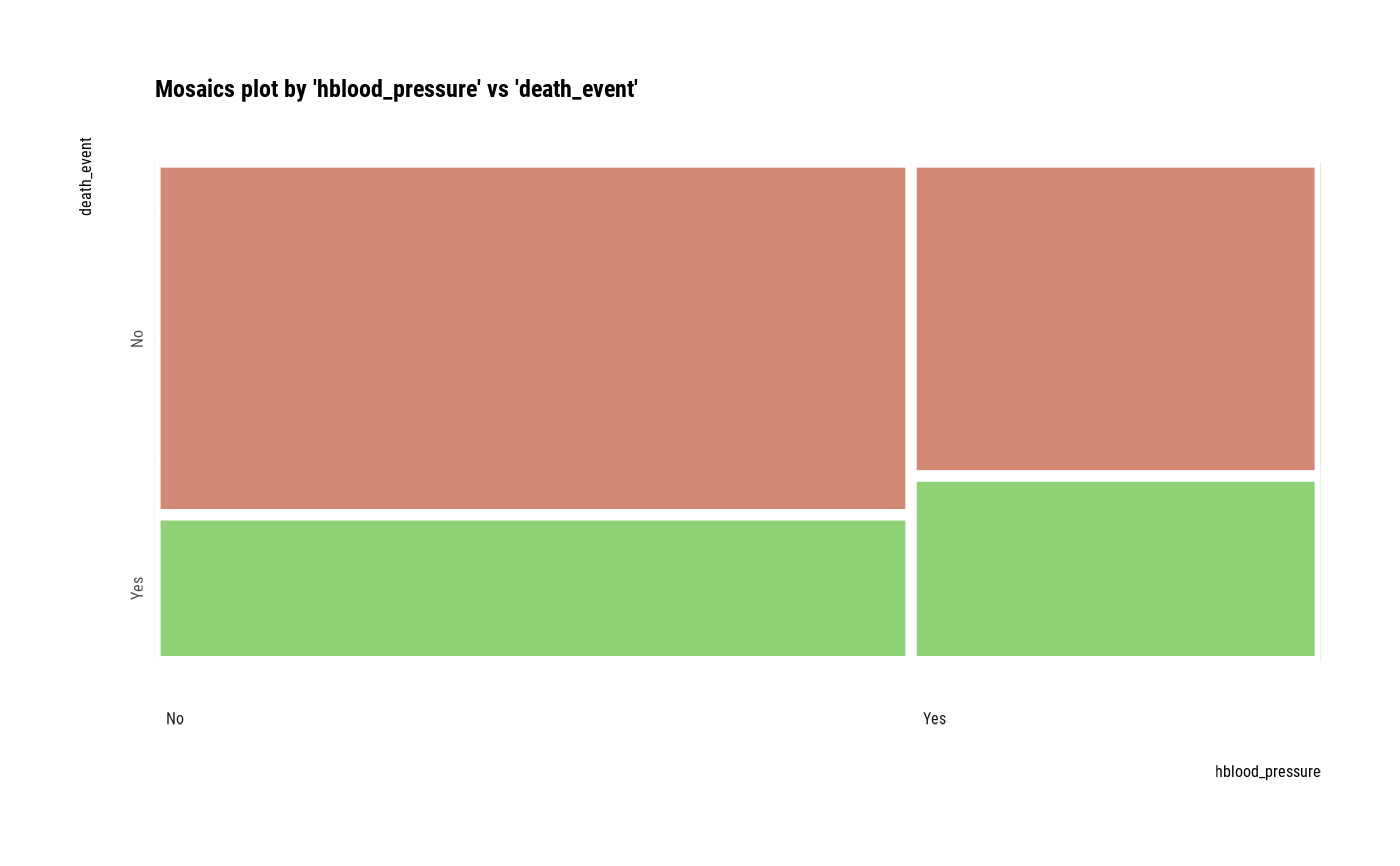
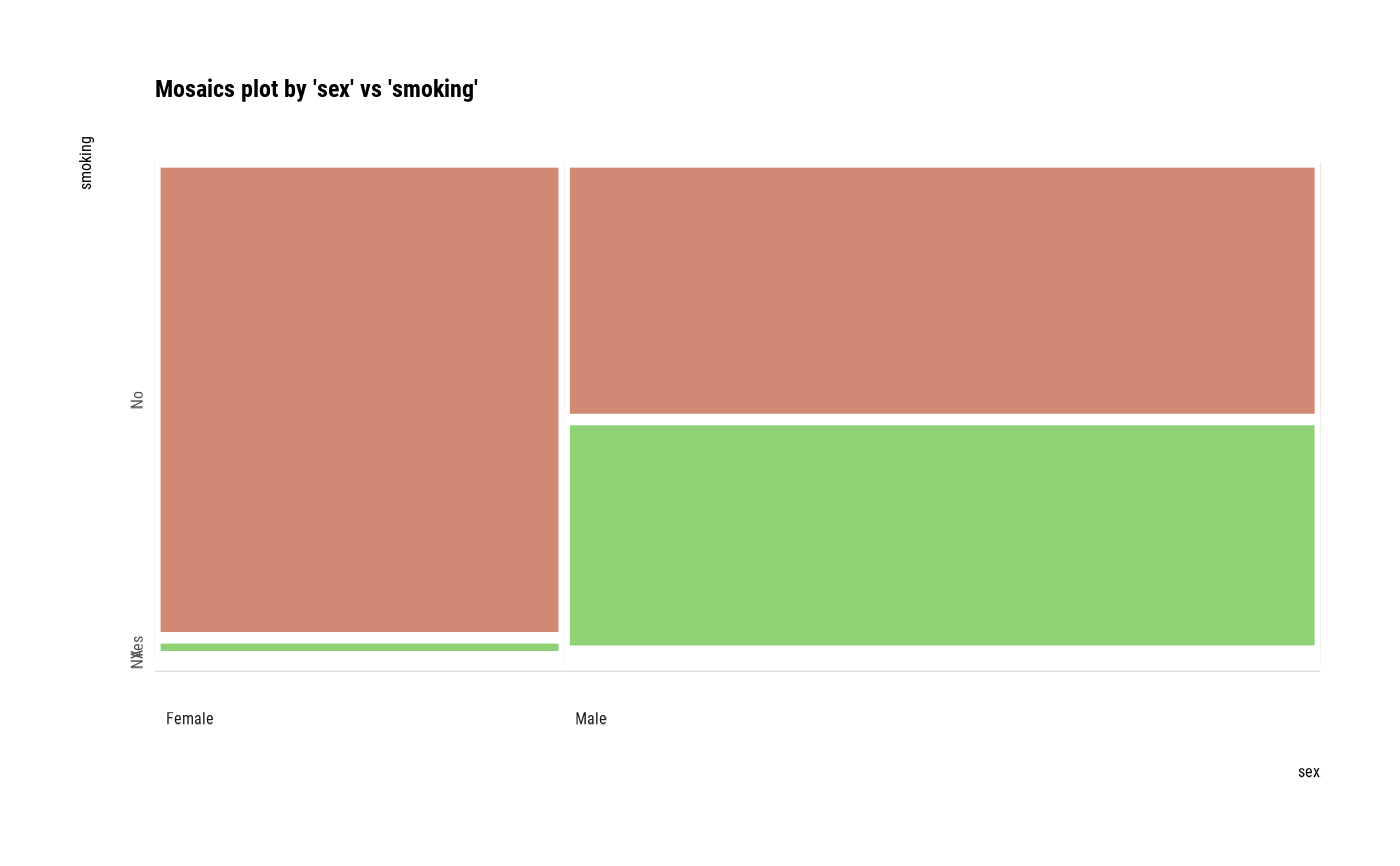
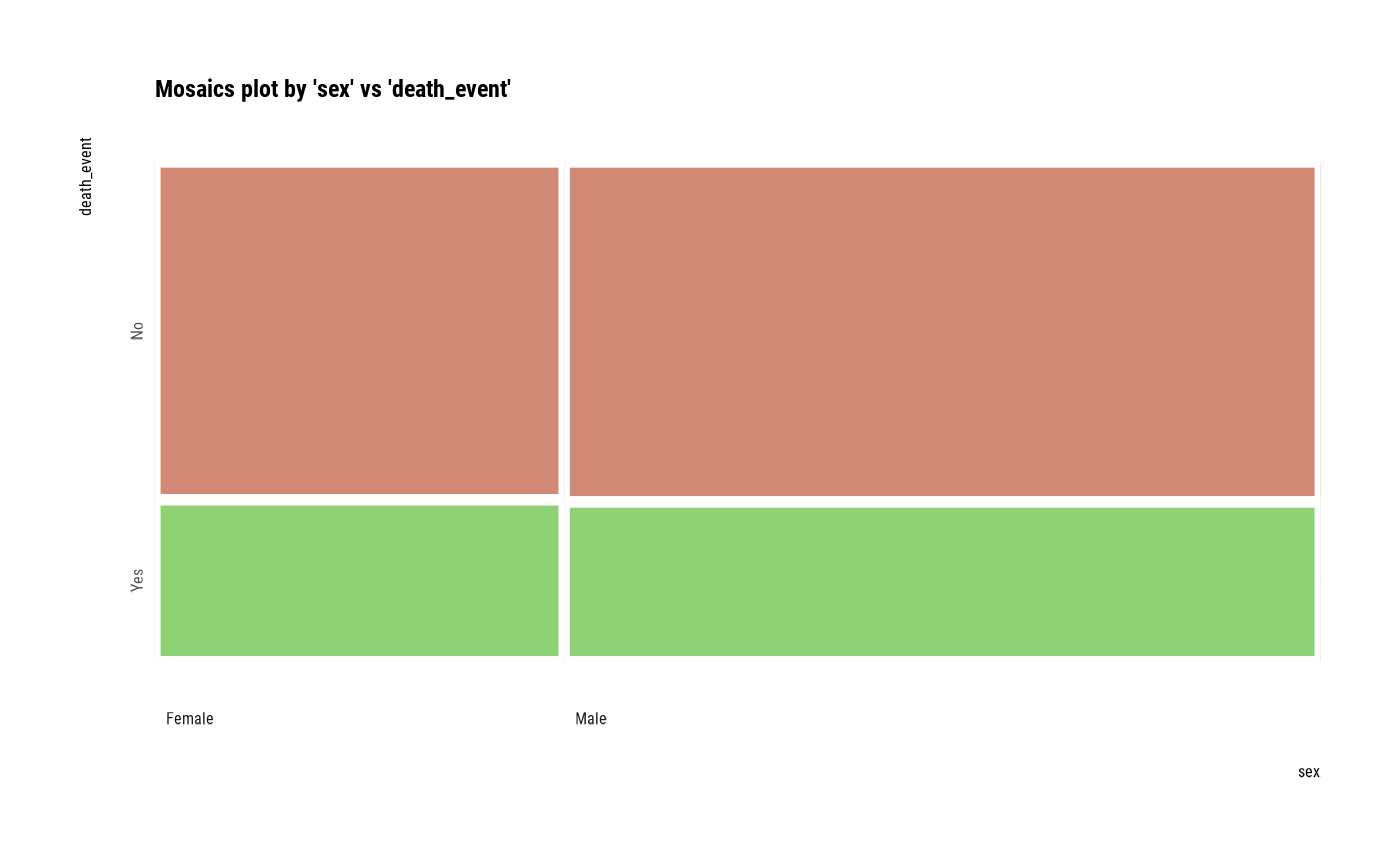
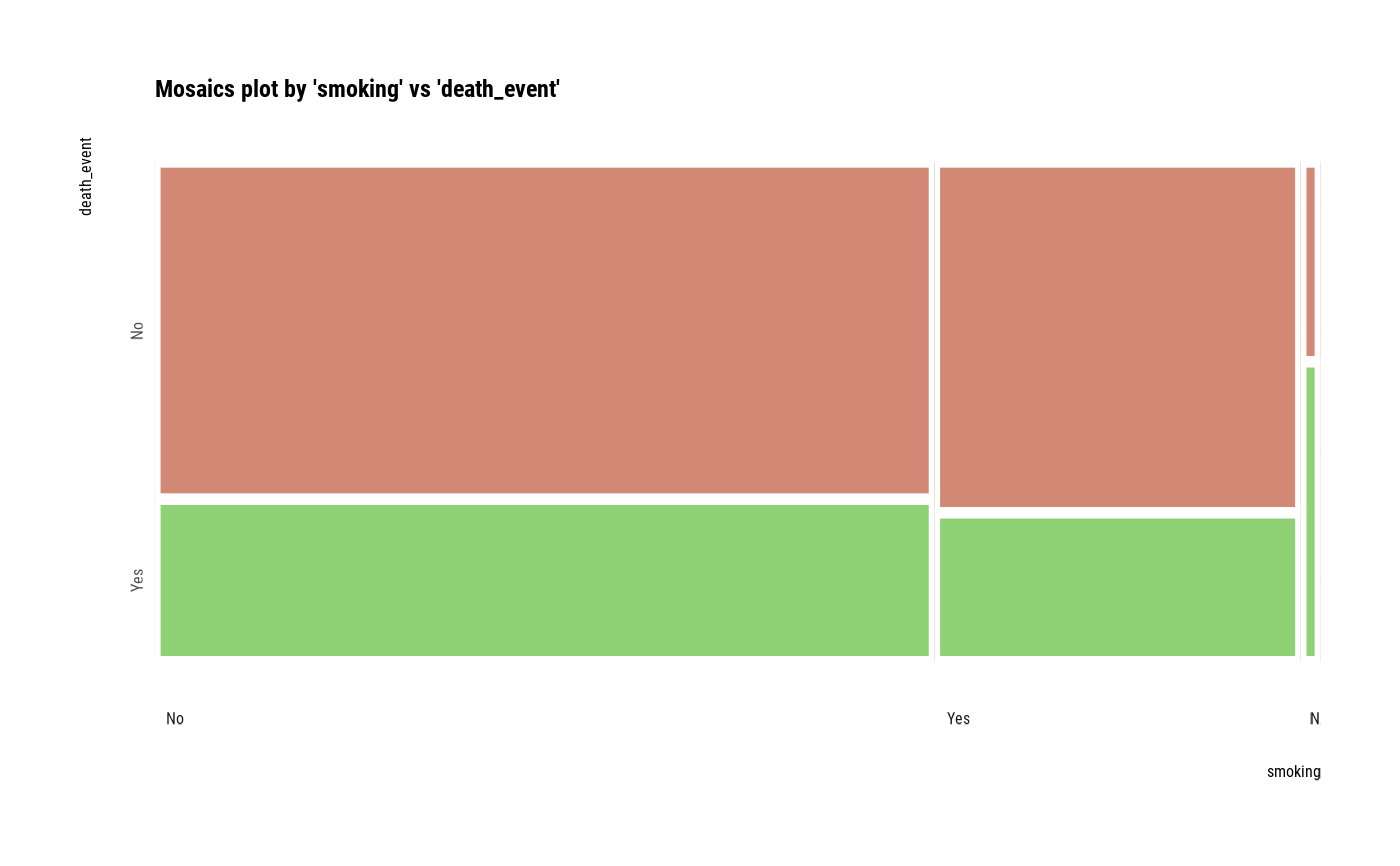
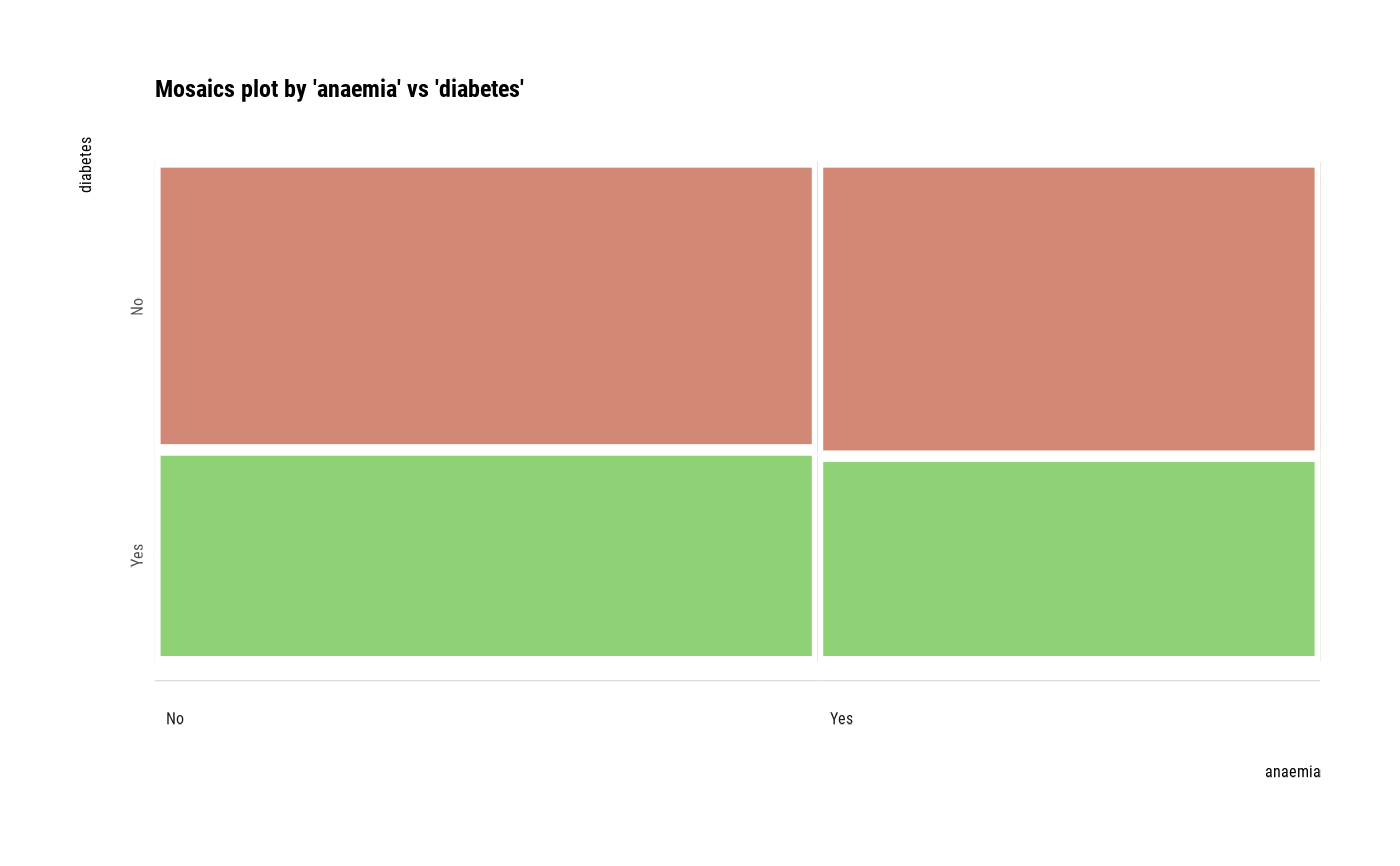
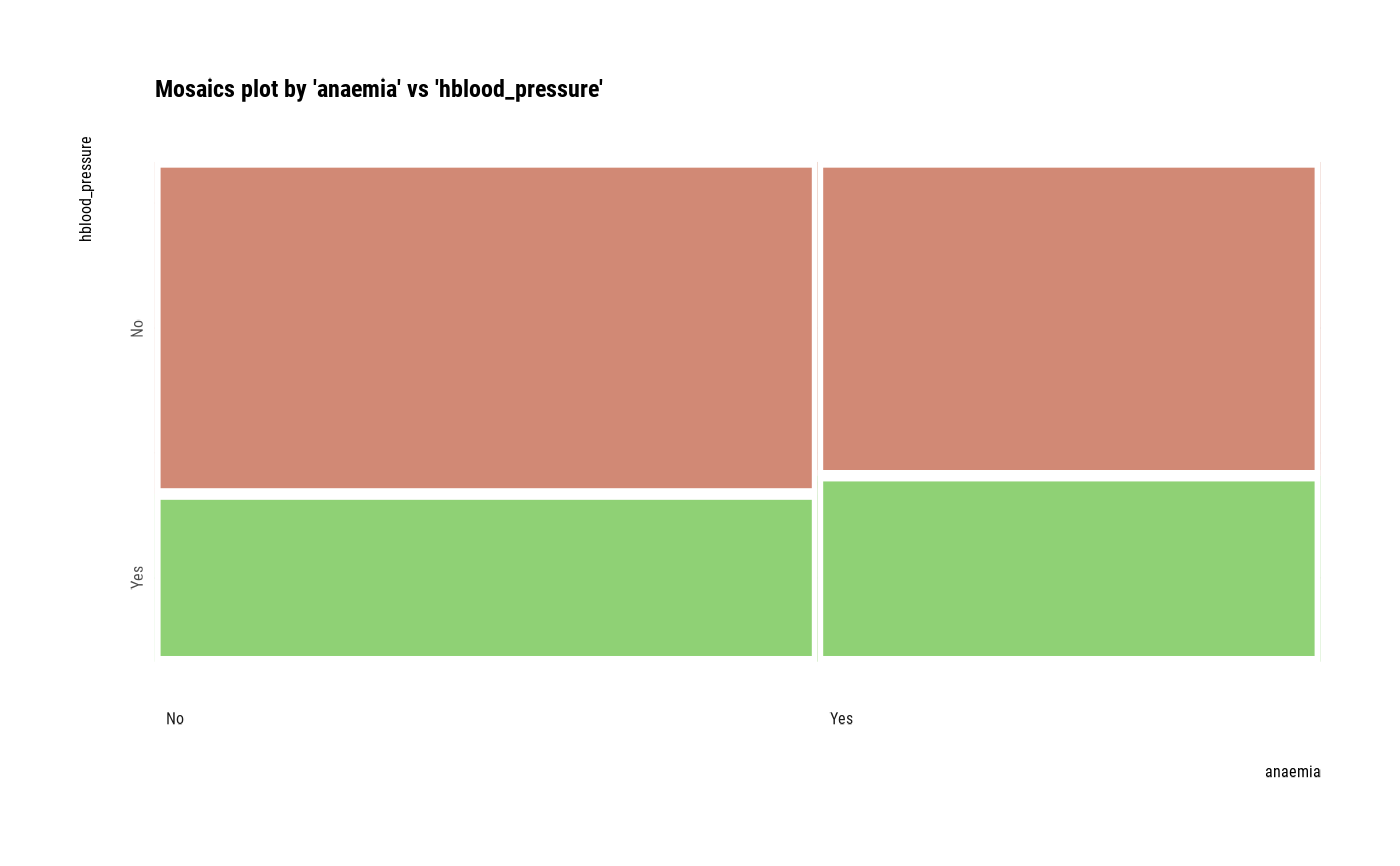
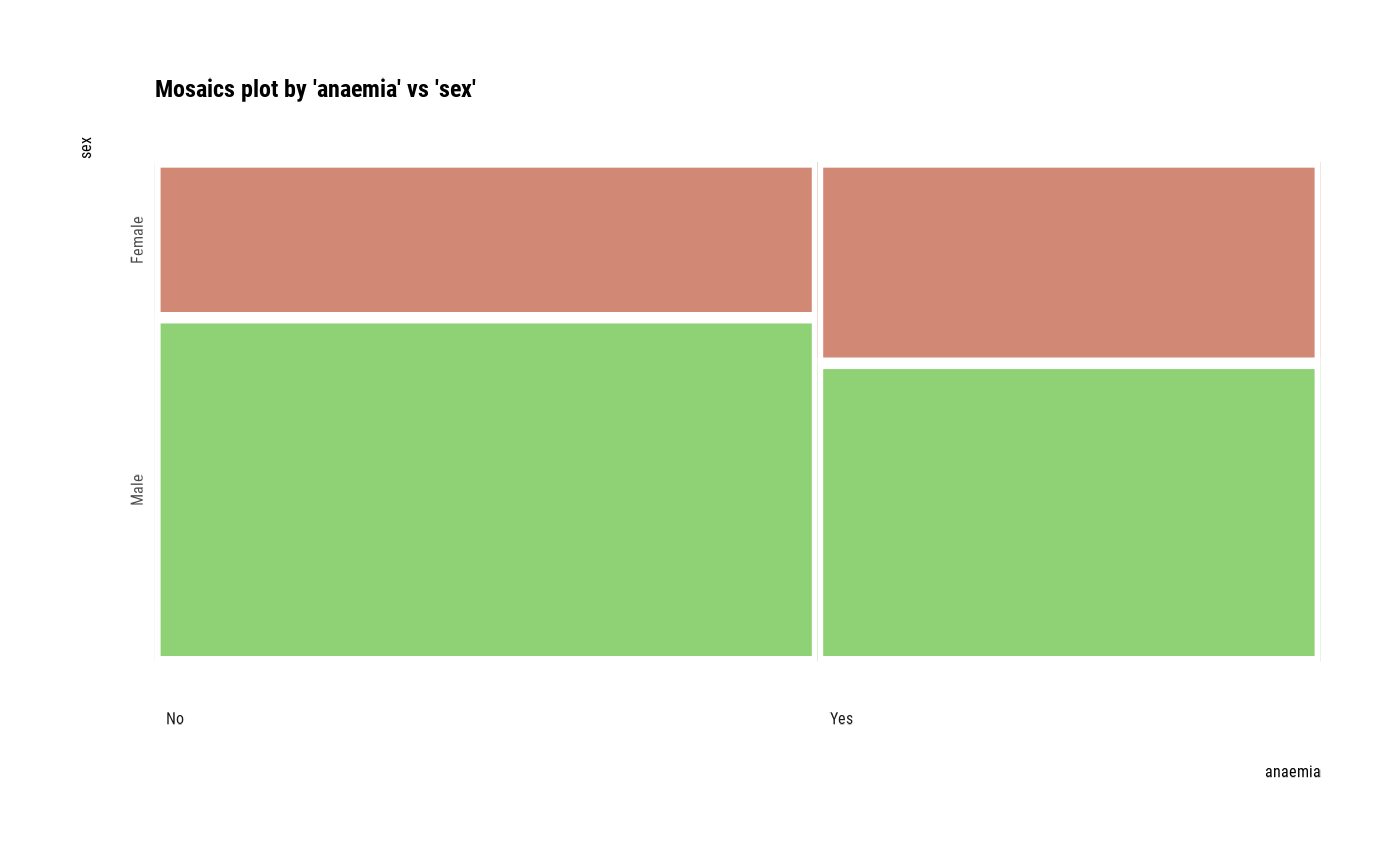
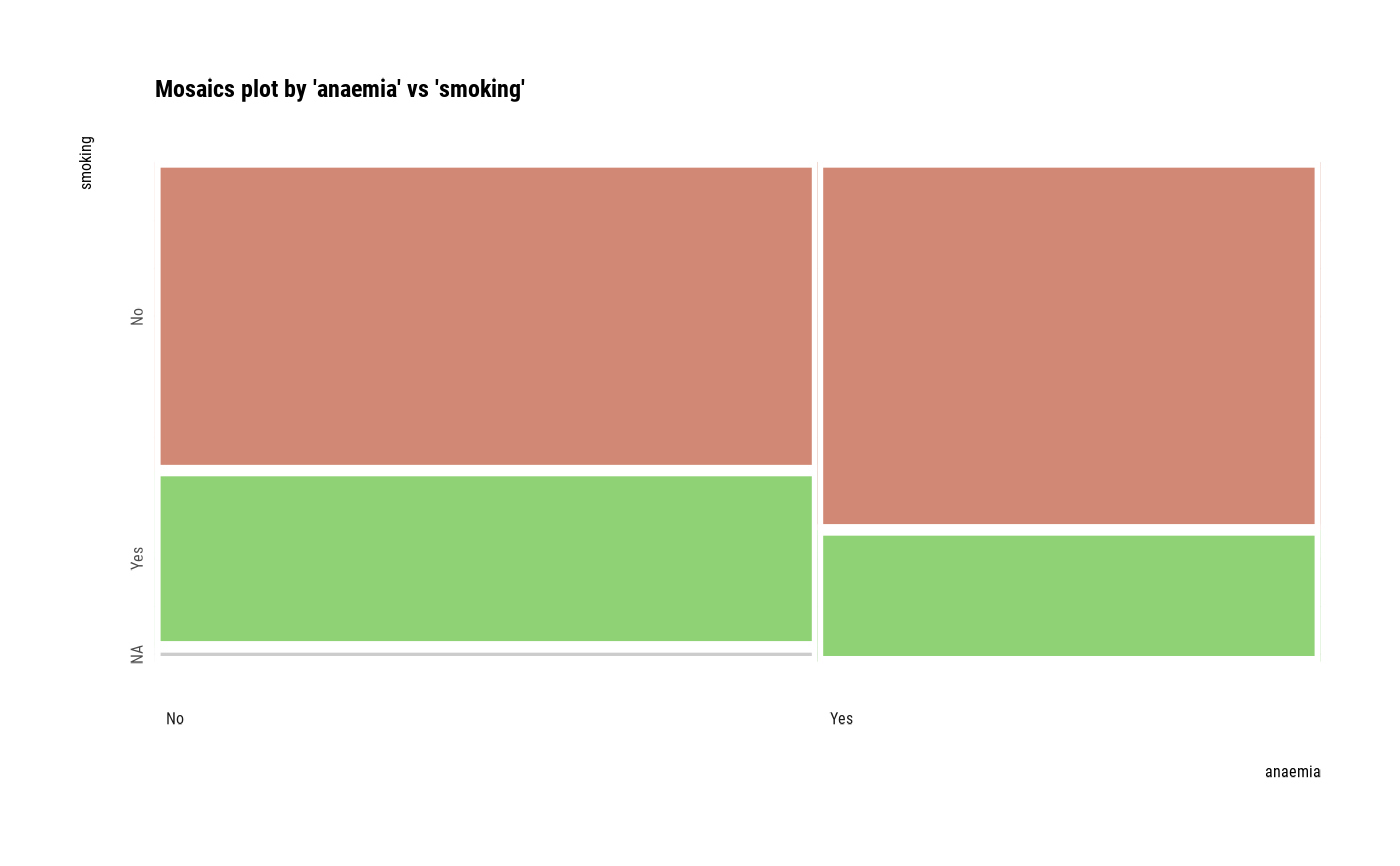
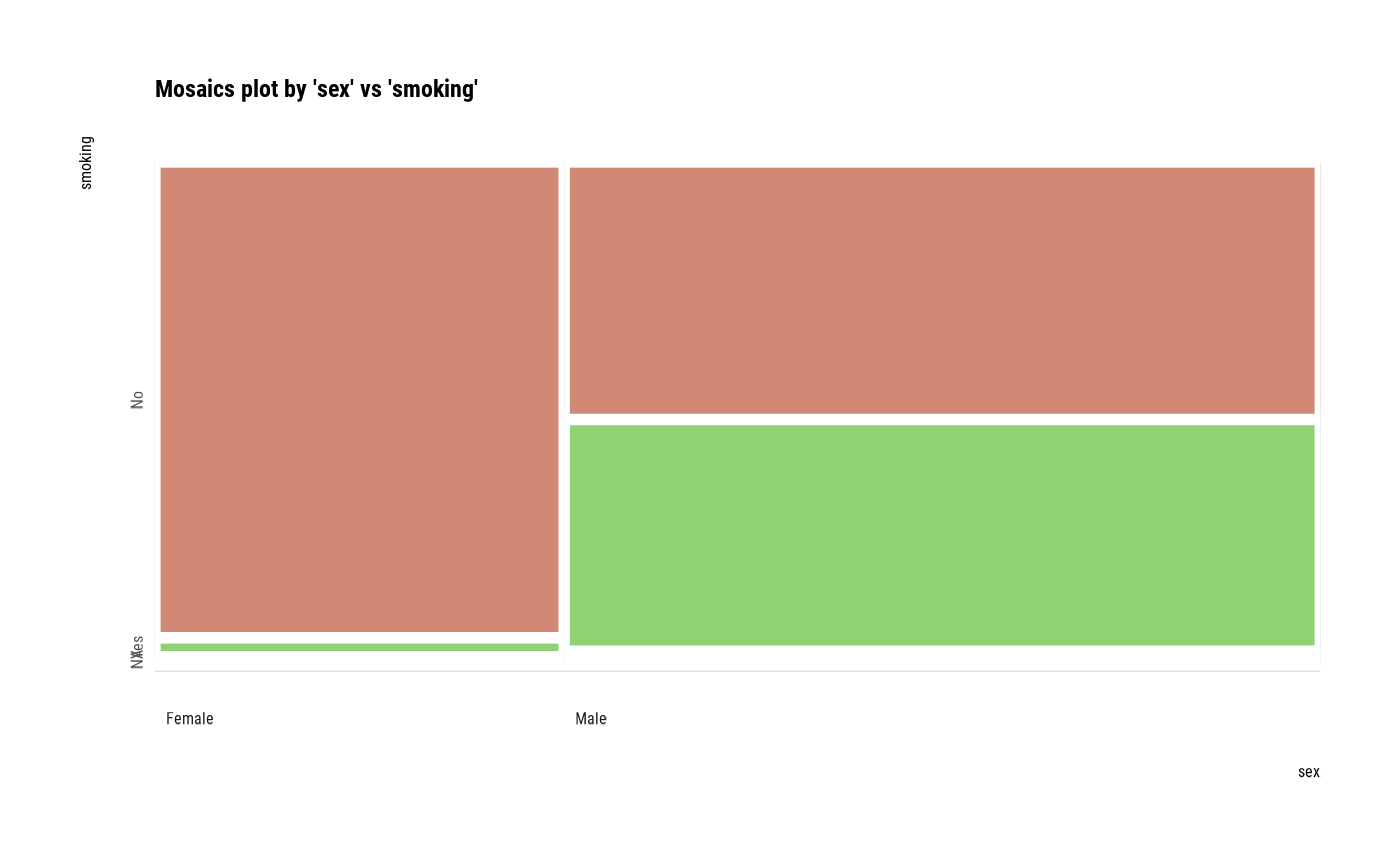
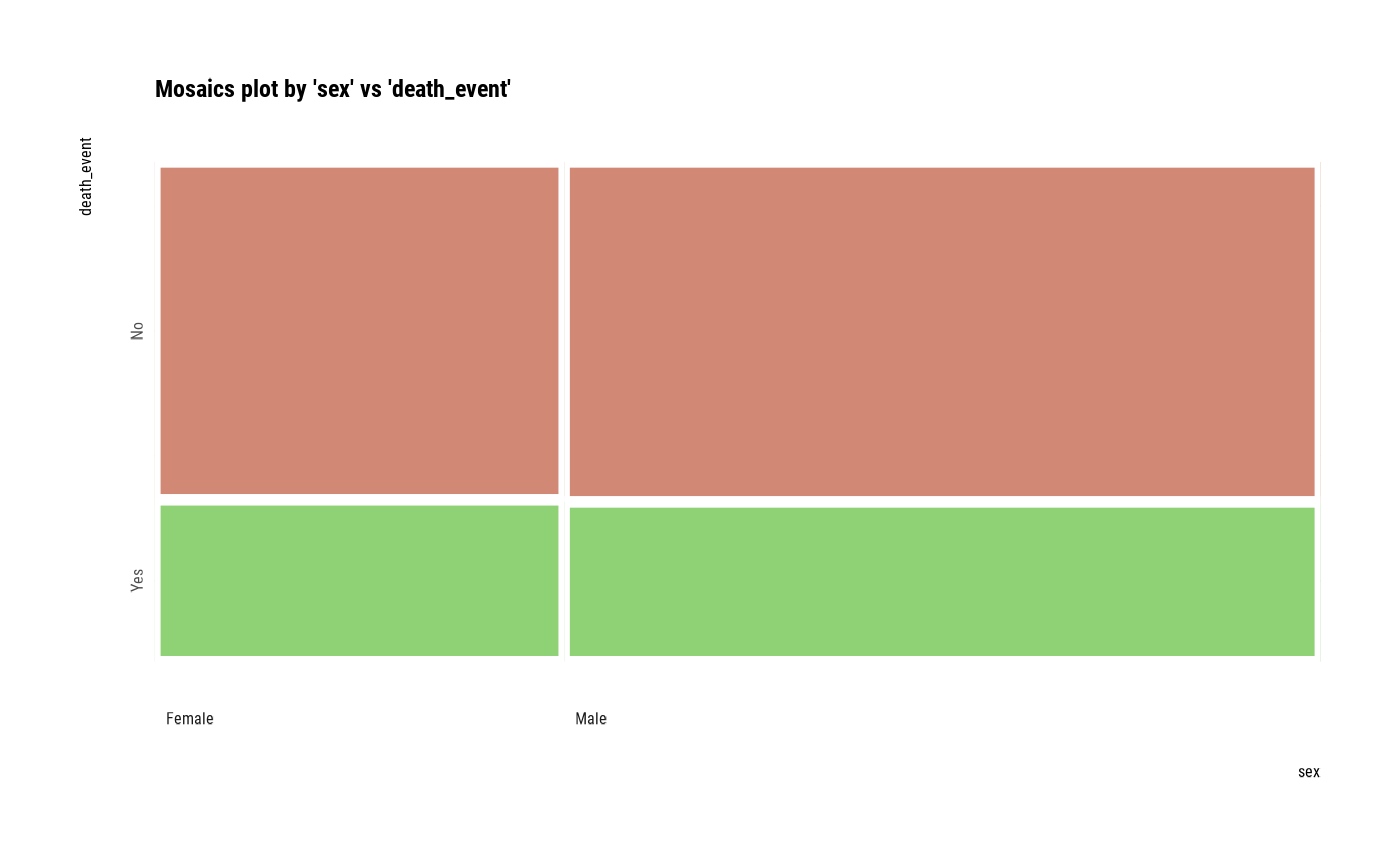
# plot all pair of variables
plot(all_var)
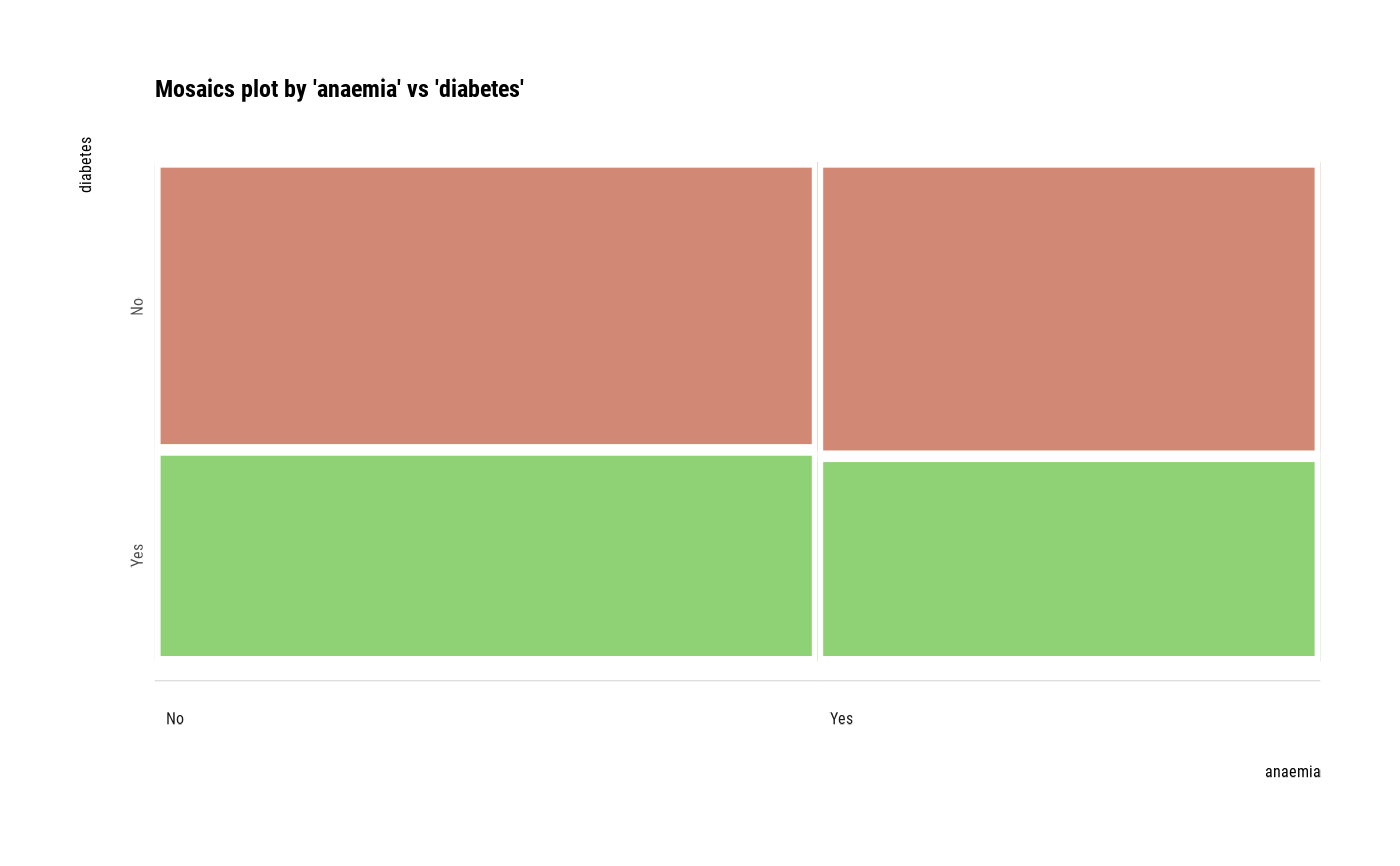
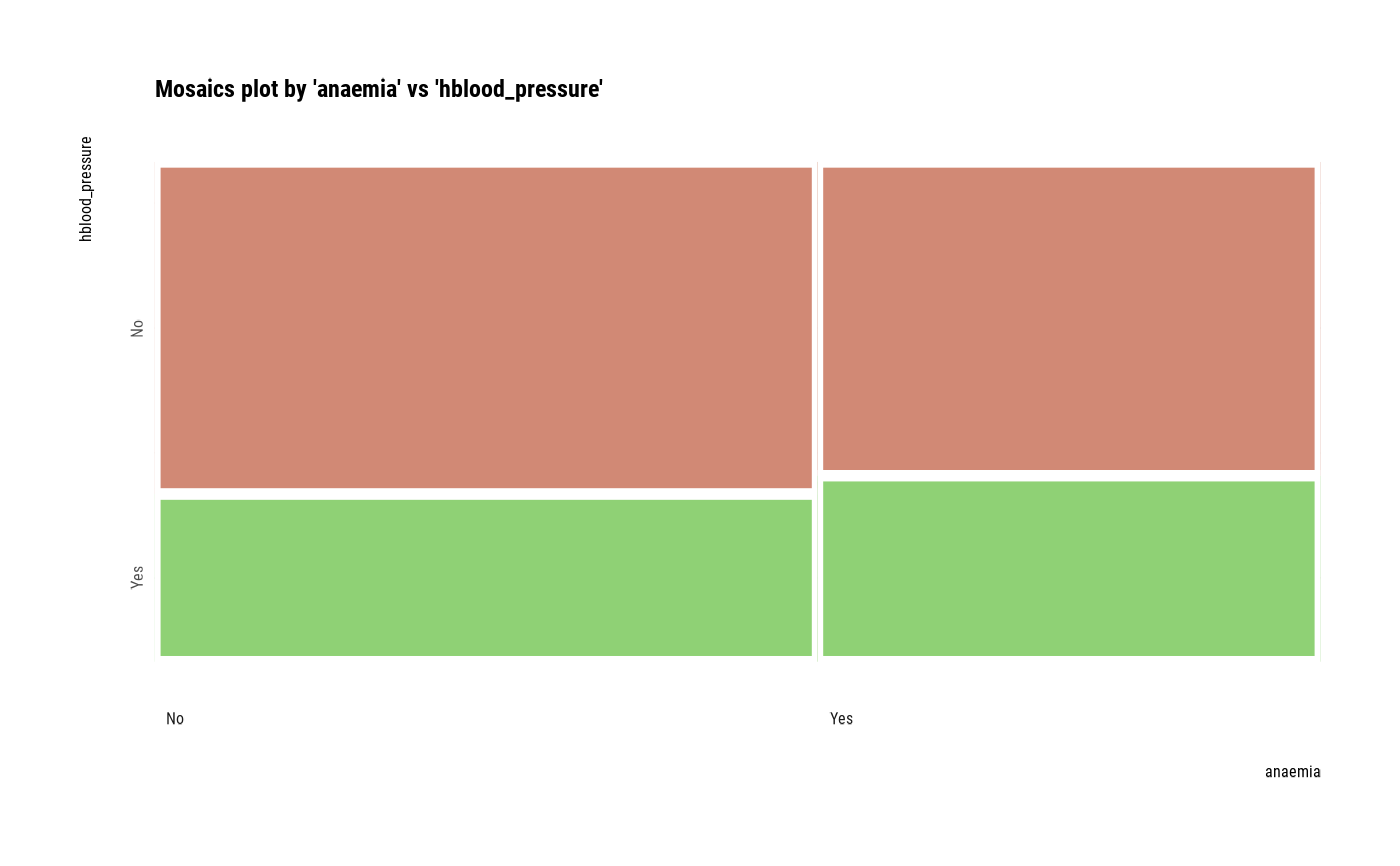
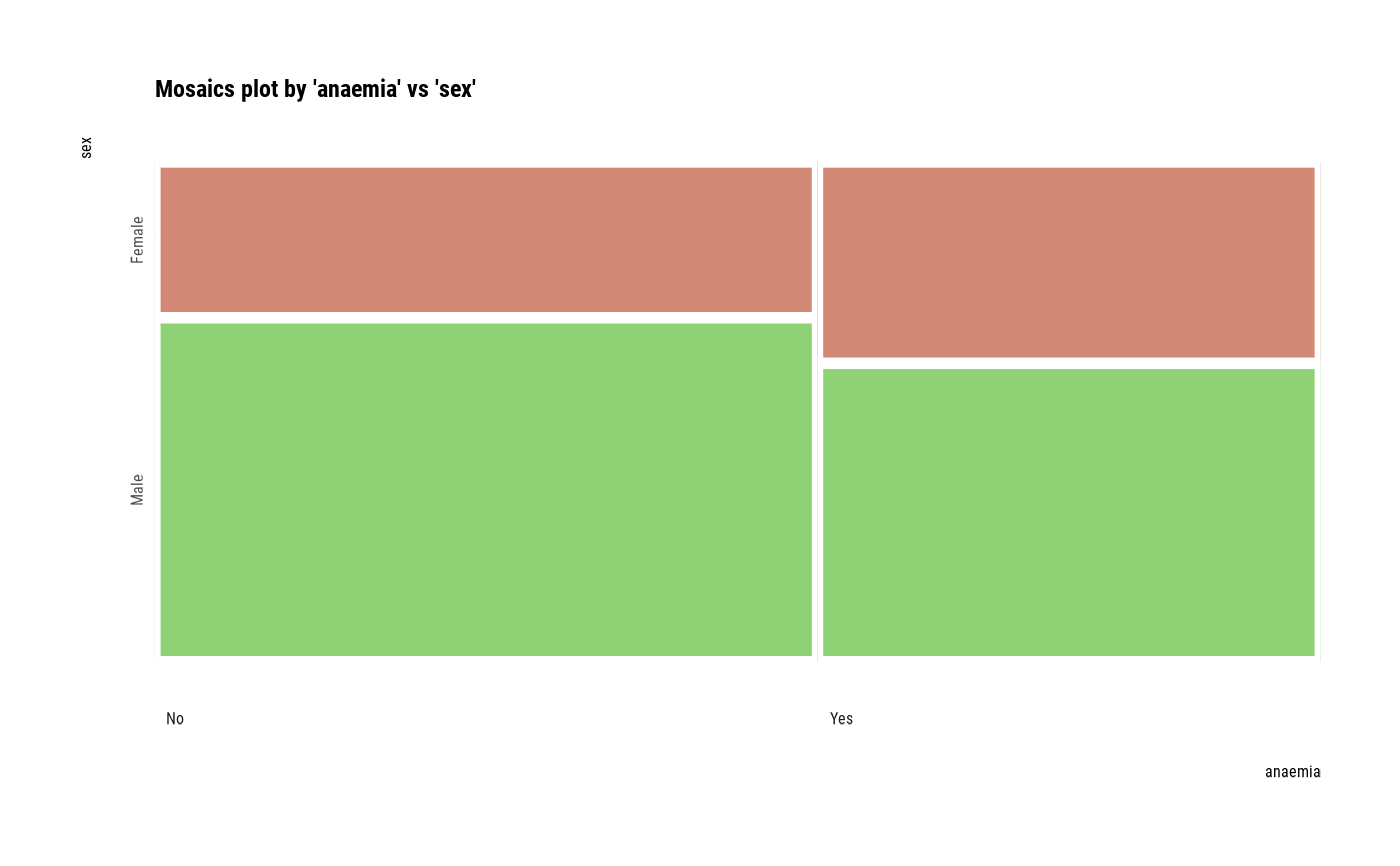
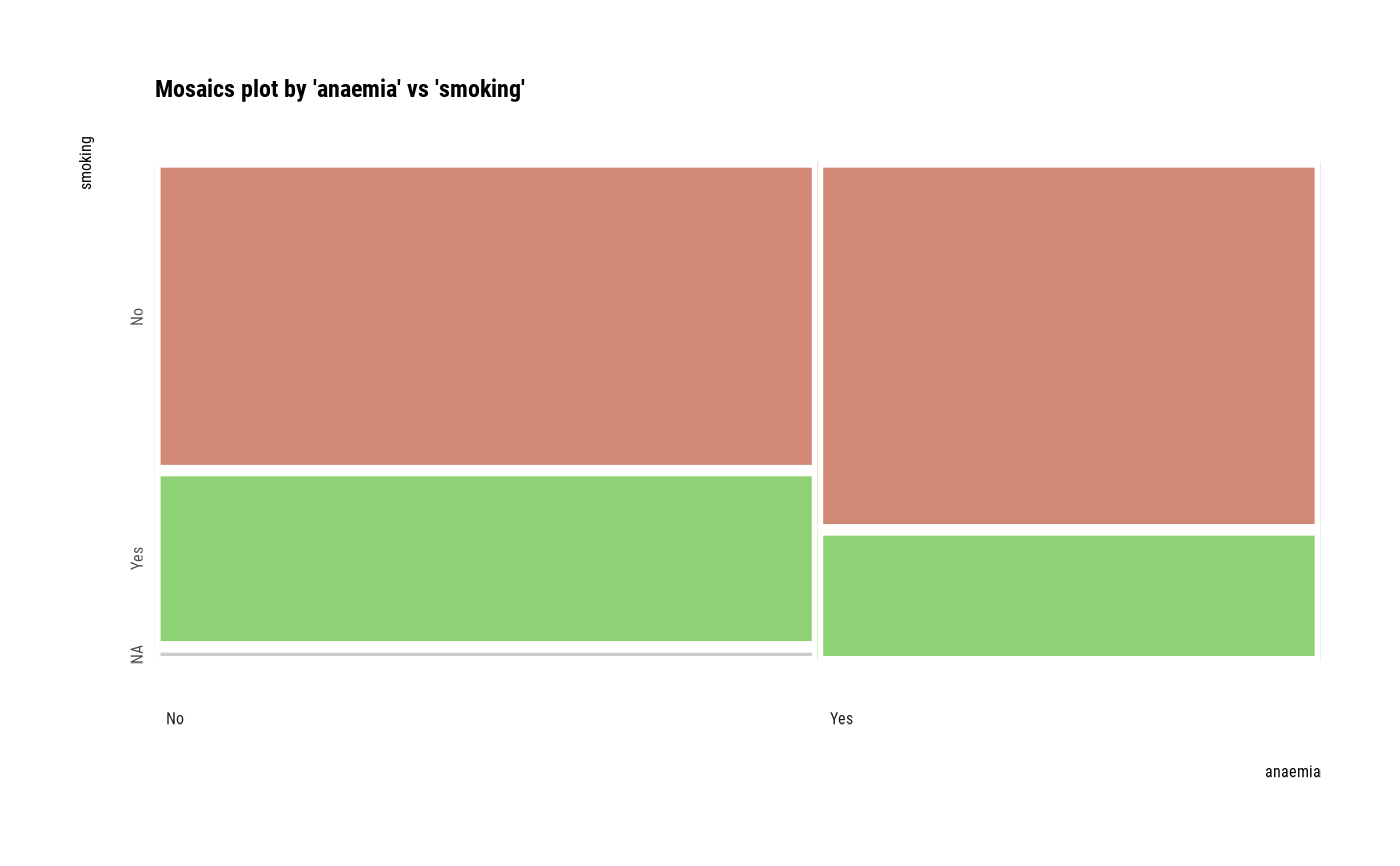
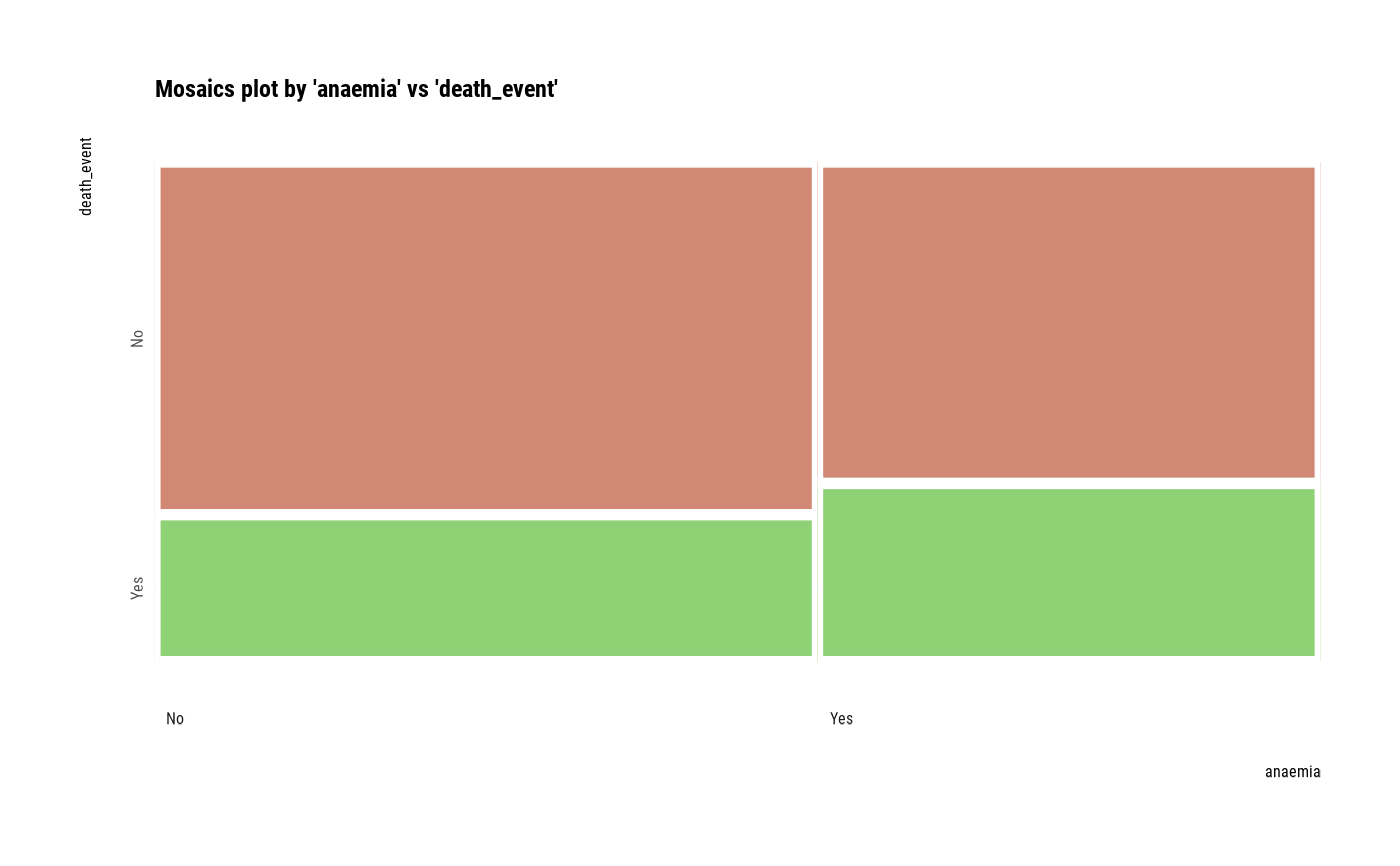
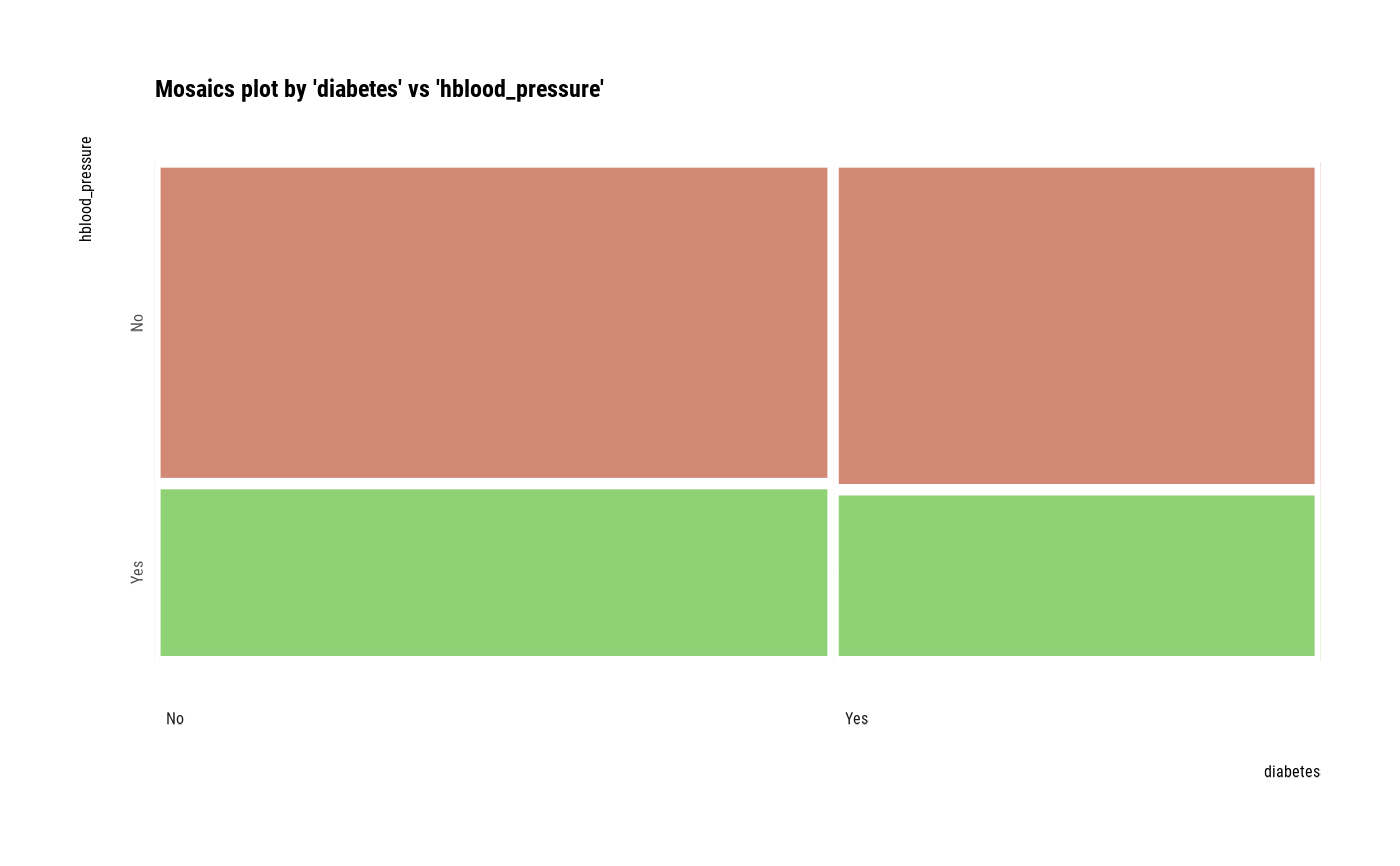
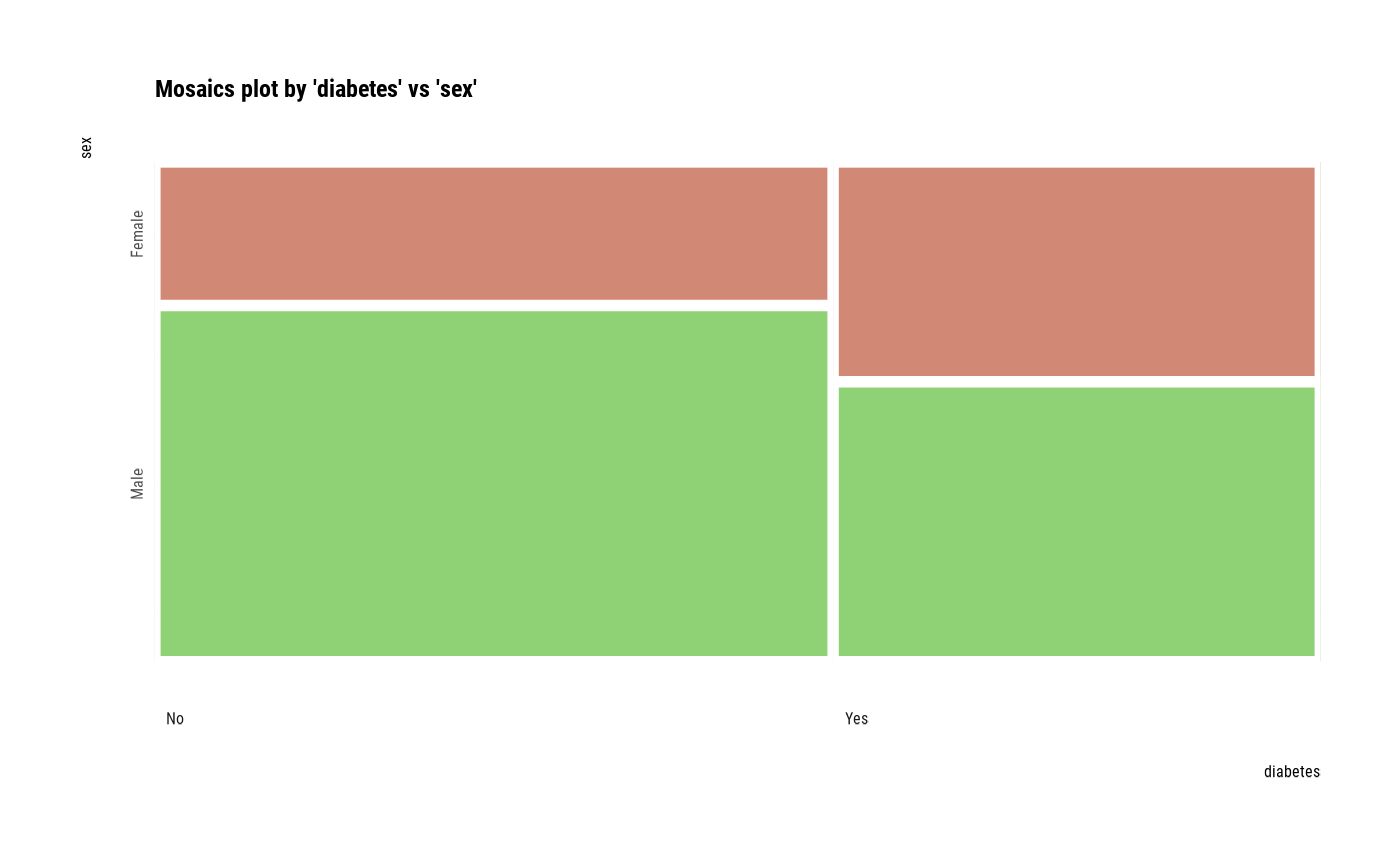
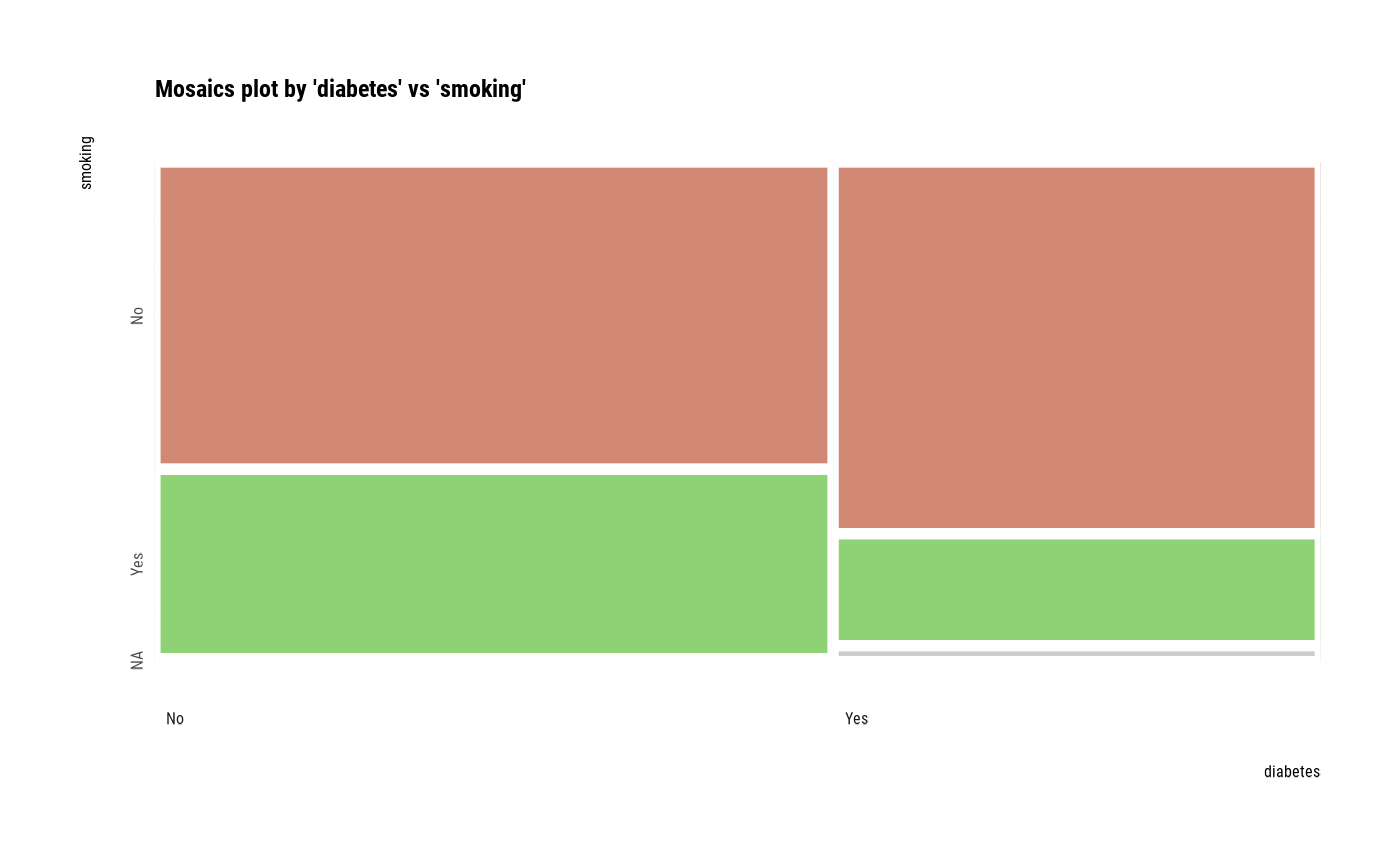
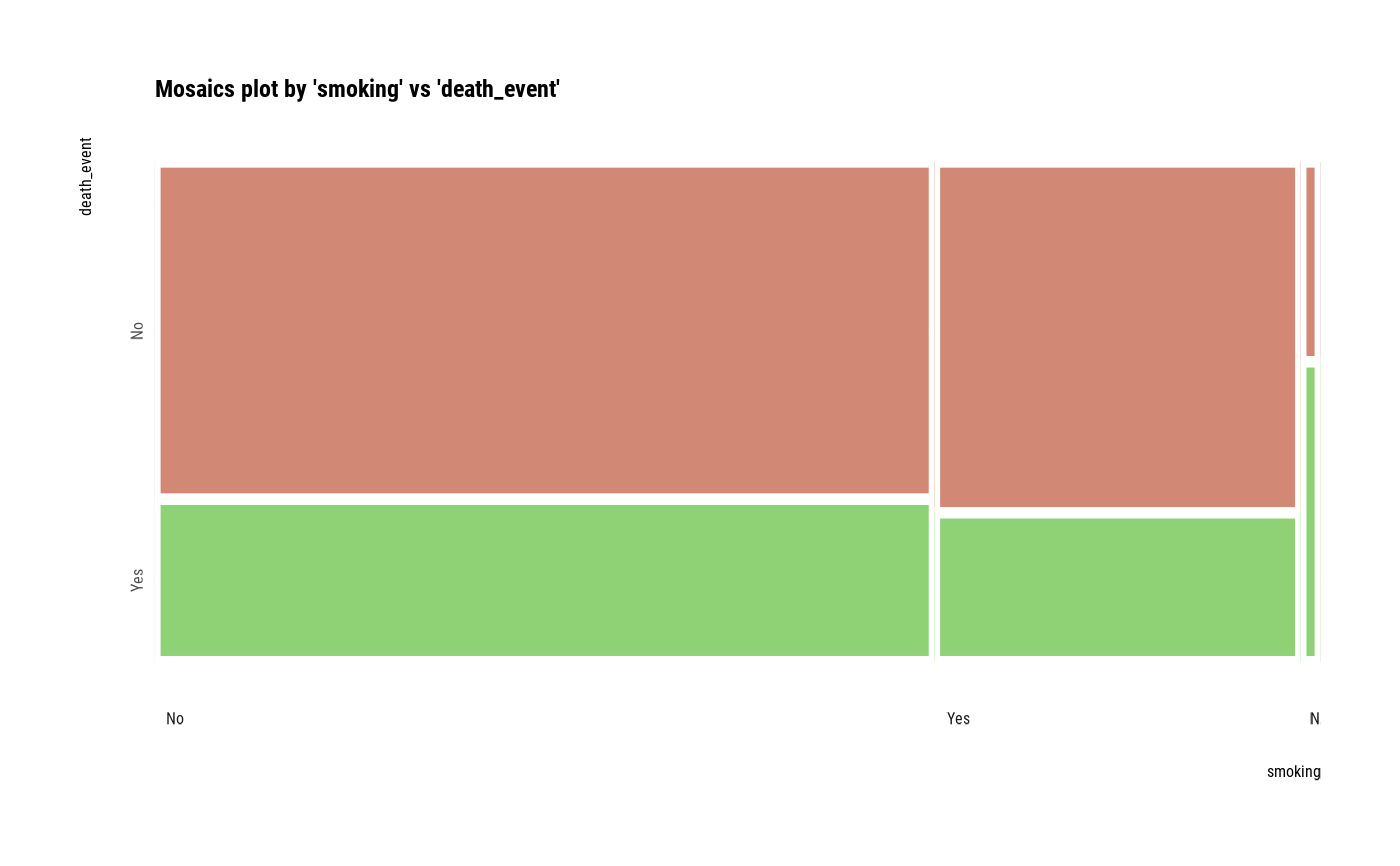
 # plot a pair of variables
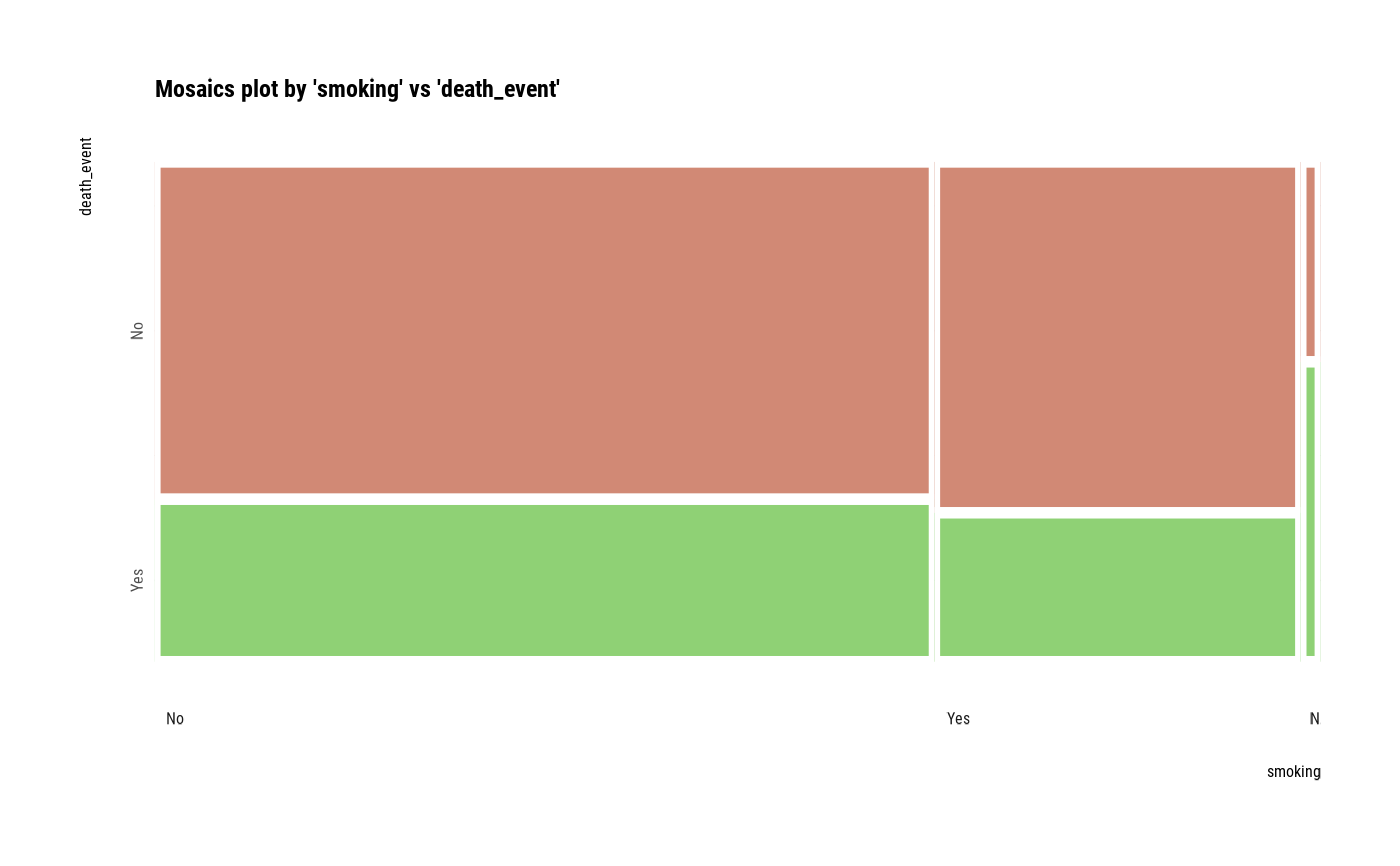
plot(two_var)
# plot a pair of variables
plot(two_var)
 # plot all pair of variables by prompt
plot(all_var, prompt = TRUE)
# plot all pair of variables by prompt
plot(all_var, prompt = TRUE)
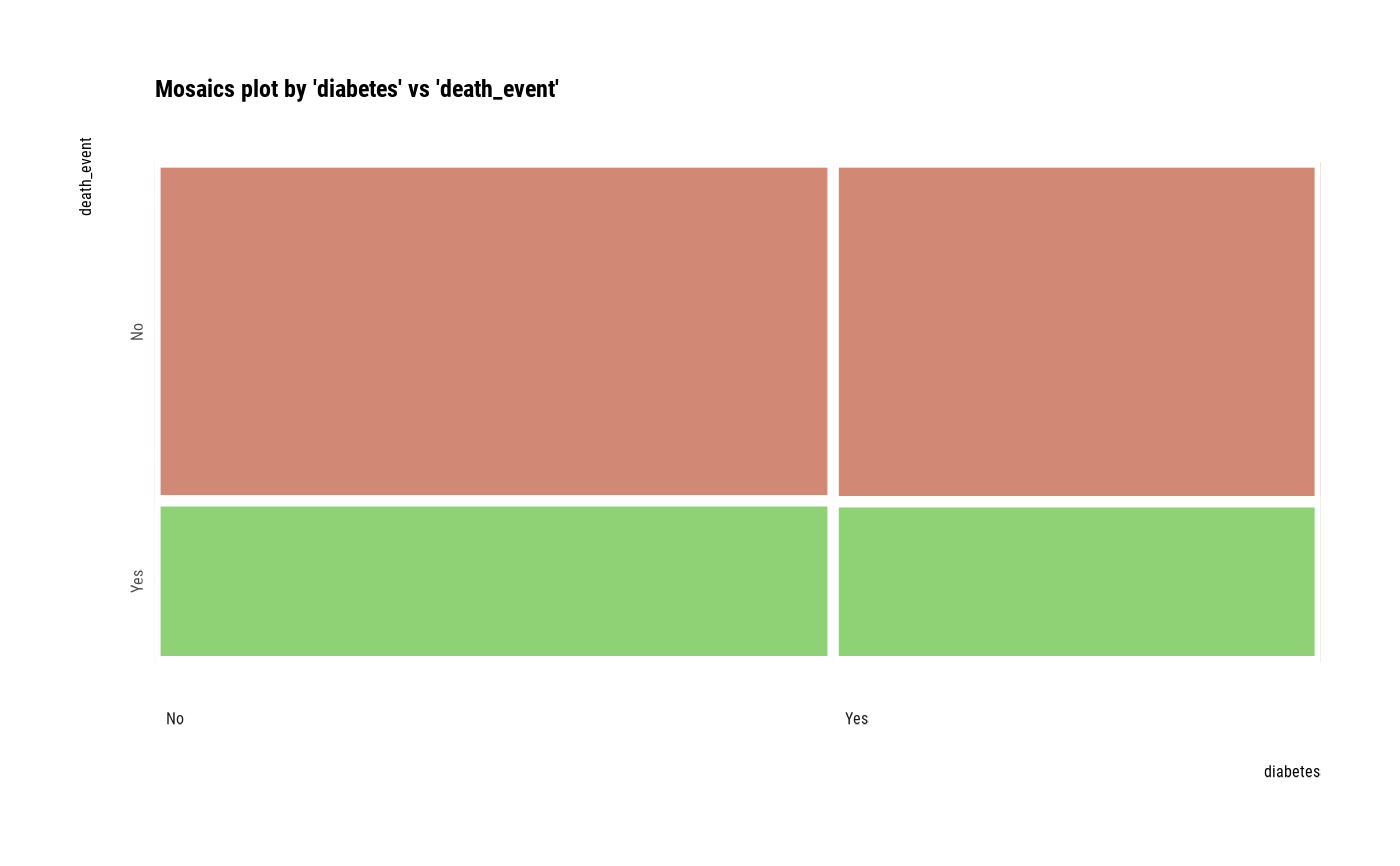
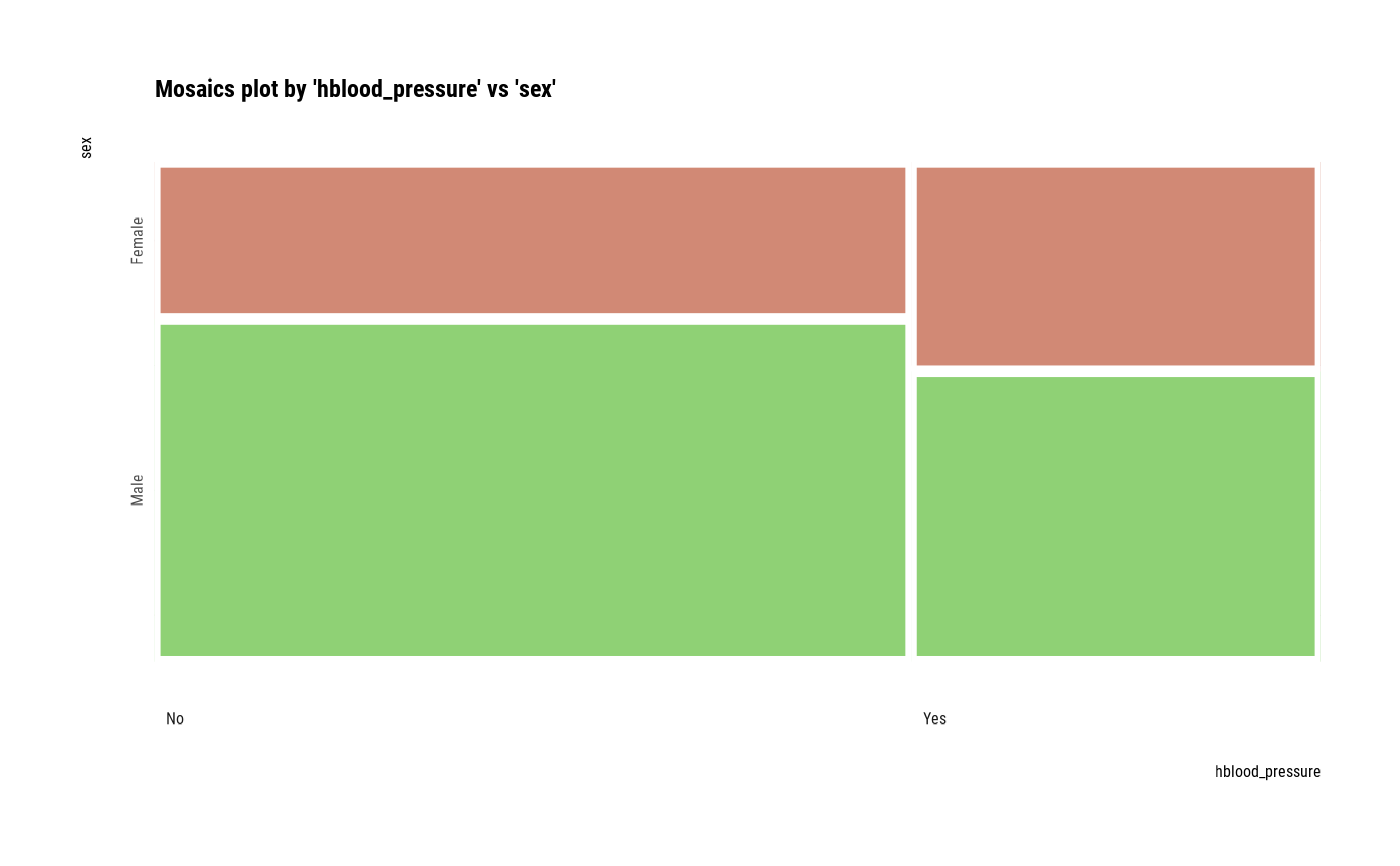
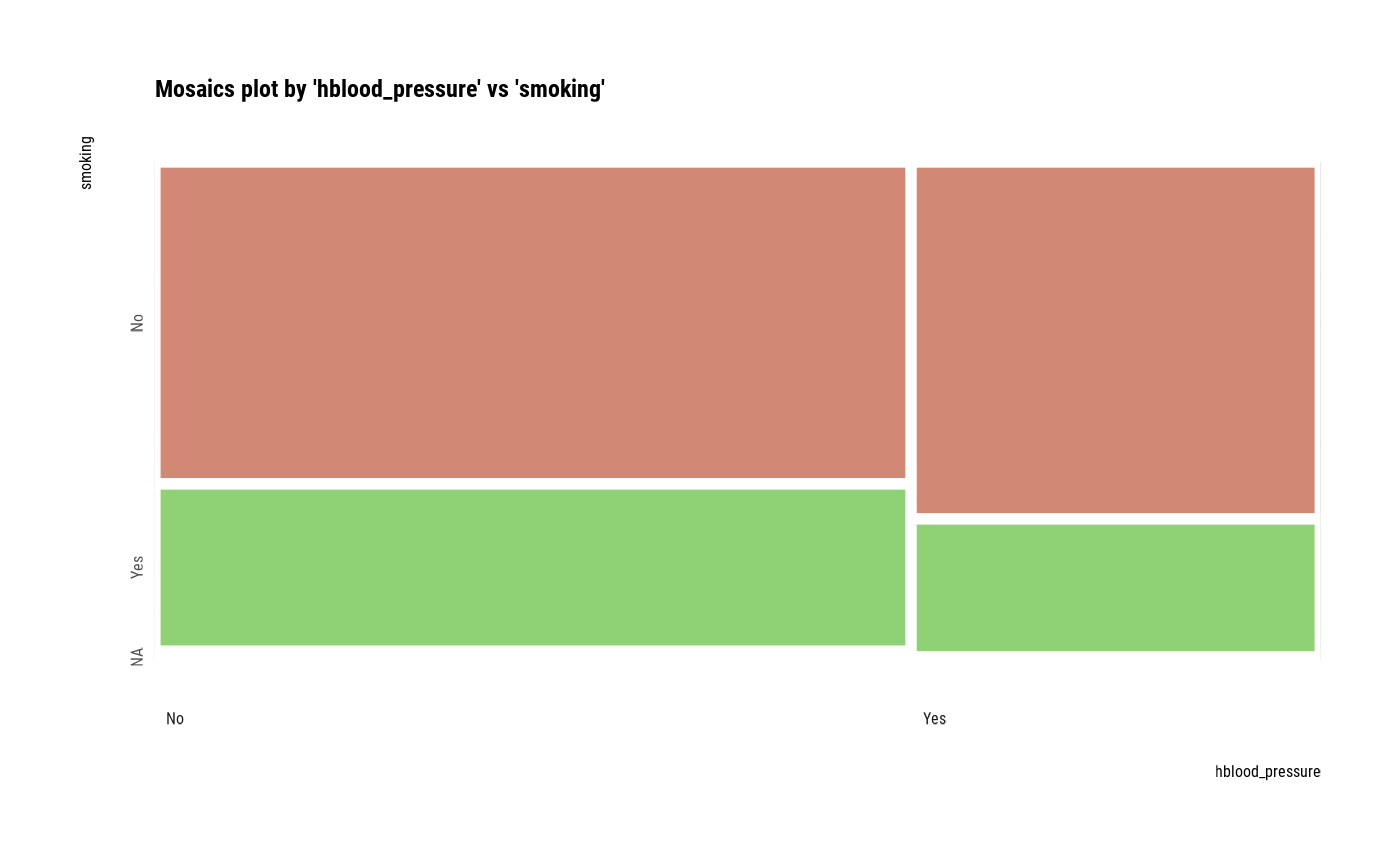
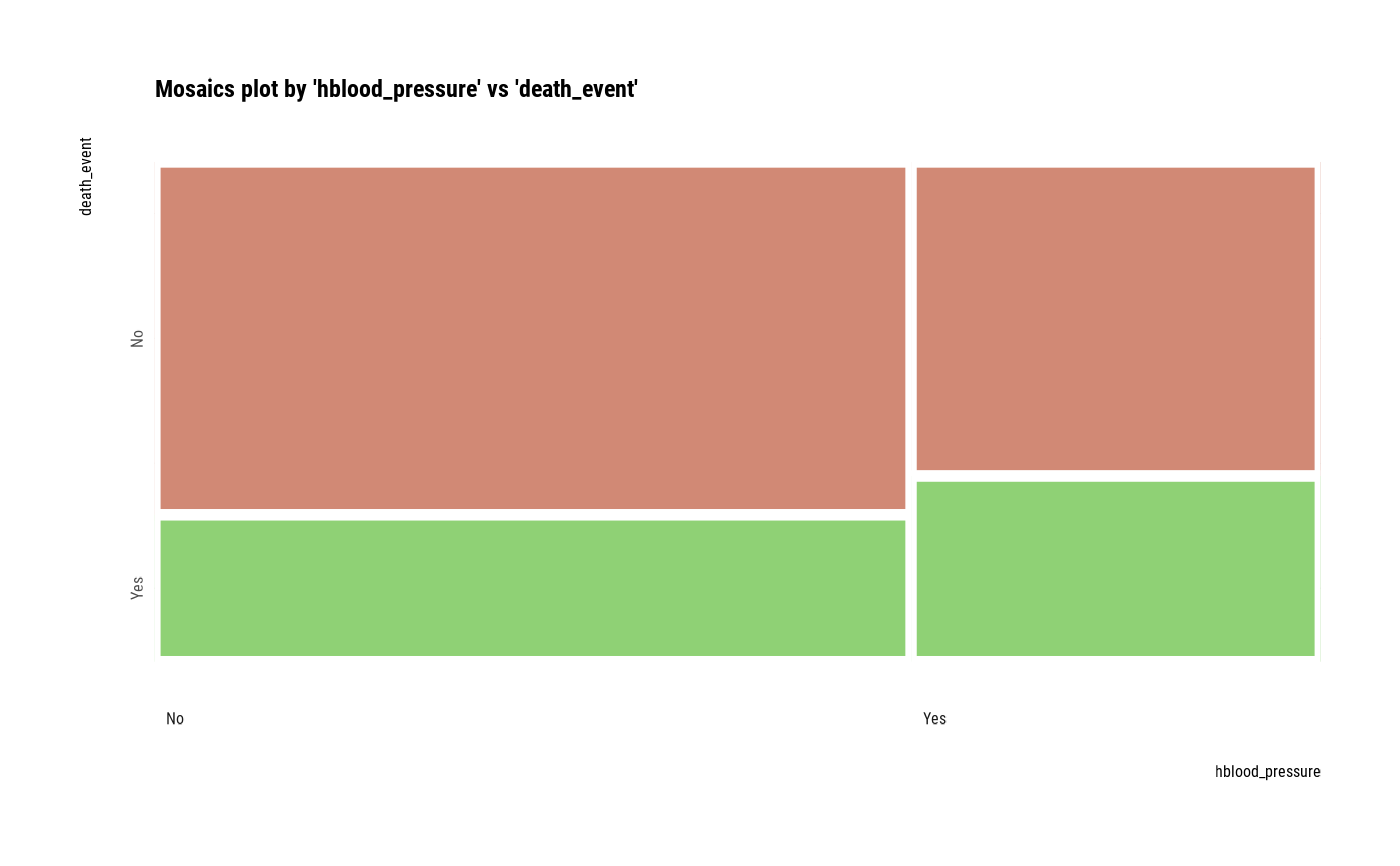
 # plot a pair of variables
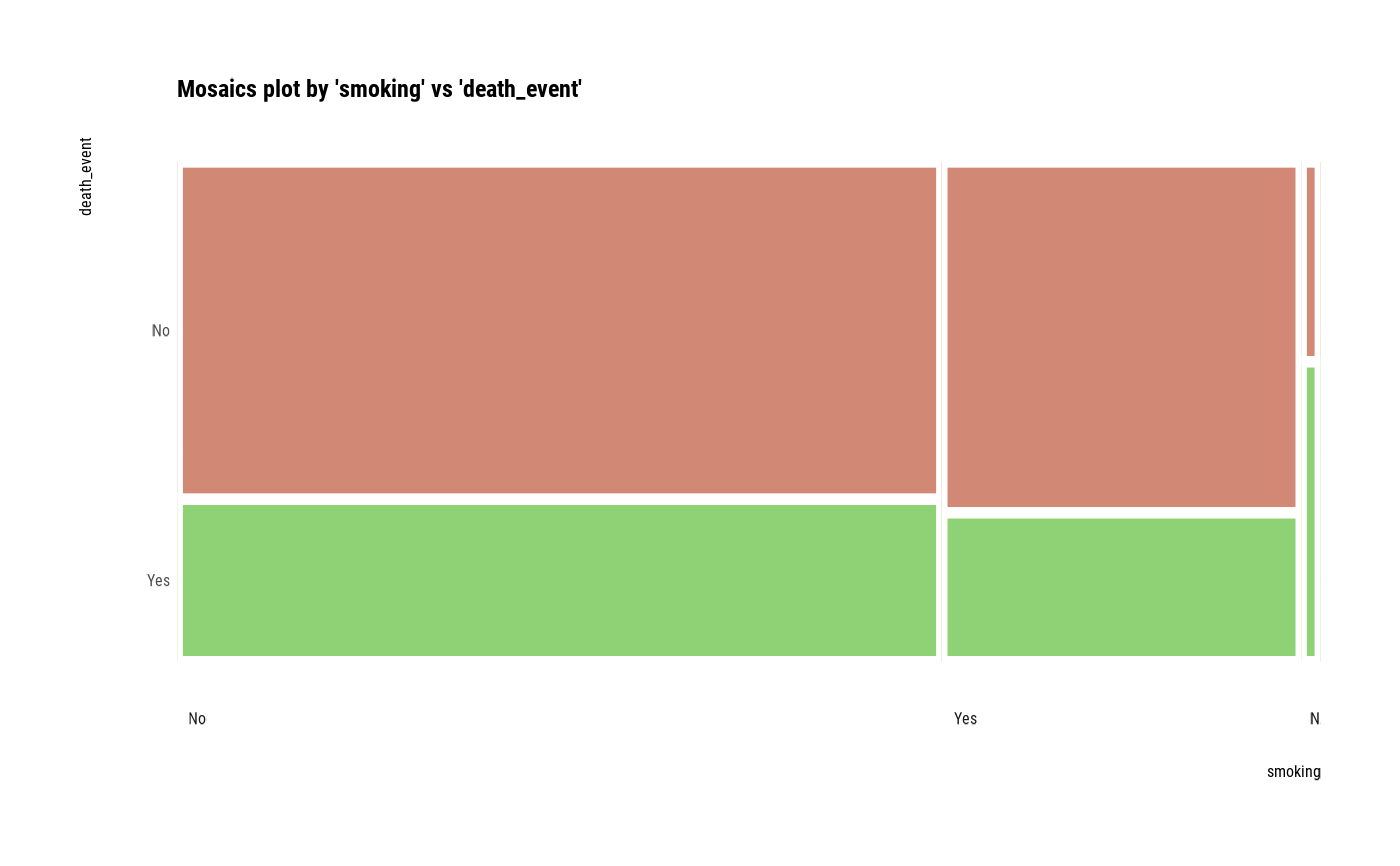
plot(two_var, las = 1)
# plot a pair of variables
plot(two_var, las = 1)
 # }
# }